Predicting essential components of signal transduction networks: a dynamic model of guard cell abscisic acid signaling
- PMID: 16968132
- PMCID: PMC1564158
- DOI: 10.1371/journal.pbio.0040312
Predicting essential components of signal transduction networks: a dynamic model of guard cell abscisic acid signaling
Abstract
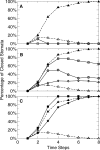
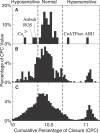
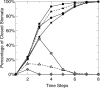
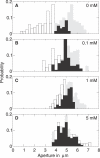
Plants both lose water and take in carbon dioxide through microscopic stomatal pores, each of which is regulated by a surrounding pair of guard cells. During drought, the plant hormone abscisic acid (ABA) inhibits stomatal opening and promotes stomatal closure, thereby promoting water conservation. Dozens of cellular components have been identified to function in ABA regulation of guard cell volume and thus of stomatal aperture, but a dynamic description is still not available for this complex process. Here we synthesize experimental results into a consistent guard cell signal transduction network for ABA-induced stomatal closure, and develop a dynamic model of this process. Our model captures the regulation of more than 40 identified network components, and accords well with previous experimental results at both the pathway and whole-cell physiological level. By simulating gene disruptions and pharmacological interventions we find that the network is robust against a significant fraction of possible perturbations. Our analysis reveals the novel predictions that the disruption of membrane depolarizability, anion efflux, actin cytoskeleton reorganization, cytosolic pH increase, the phosphatidic acid pathway, or K(+) efflux through slowly activating K(+) channels at the plasma membrane lead to the strongest reduction in ABA responsiveness. Initial experimental analysis assessing ABA-induced stomatal closure in the presence of cytosolic pH clamp imposed by the weak acid butyrate is consistent with model prediction. Simulations of stomatal response as derived from our model provide an efficient tool for the identification of candidate manipulations that have the best chance of conferring increased drought stress tolerance and for the prioritization of future wet bench analyses. Our method can be readily applied to other biological signaling networks to identify key regulatory components in systems where quantitative information is limited.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures
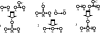
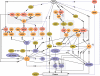
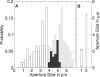
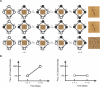
Comment in
-
When evidence is scant, mathematical modeling offers a roadmap for discovery.PLoS Biol. 2006 Oct;4(10):e323. doi: 10.1371/journal.pbio.0040323. Epub 2006 Sep 12. PLoS Biol. 2006. PMID: 20076468 Free PMC article. No abstract available.
Similar articles
-
Mitochondrial pyruvate carrier 1 mediates abscisic acid-regulated stomatal closure and the drought response by affecting cellular pyruvate content in Arabidopsis thaliana.BMC Plant Biol. 2017 Nov 22;17(1):217. doi: 10.1186/s12870-017-1175-3. BMC Plant Biol. 2017. PMID: 29166881 Free PMC article.
-
Identification of a novel stomatal opening chemical, PP242, that inhibits early abscisic acid signal transduction in guard cells.Plant Cell Physiol. 2025 Jul 24;66(6):854-865. doi: 10.1093/pcp/pcaf013. Plant Cell Physiol. 2025. PMID: 39882944 Free PMC article.
-
F-box protein DOR functions as a novel inhibitory factor for abscisic acid-induced stomatal closure under drought stress in Arabidopsis,Plant Physiol. 2008 Dec;148(4):2121-33. doi: 10.1104/pp.108.126912. Epub 2008 Oct 3. Plant Physiol. 2008. PMID: 18835996 Free PMC article.
-
Signal transduction and ion channels in guard cells.Philos Trans R Soc Lond B Biol Sci. 1998 Sep 29;353(1374):1475-88. doi: 10.1098/rstb.1998.0303. Philos Trans R Soc Lond B Biol Sci. 1998. PMID: 9800209 Free PMC article. Review.
-
Signaling Transduction of ABA, ROS, and Ca2+ in Plant Stomatal Closure in Response to Drought.Int J Mol Sci. 2022 Nov 26;23(23):14824. doi: 10.3390/ijms232314824. Int J Mol Sci. 2022. PMID: 36499153 Free PMC article. Review.
Cited by
-
TaRac6 Is a Potential Susceptibility Factor by Regulating the ROS Burst Negatively in the Wheat-Puccinia striiformis f. sp. tritici Interaction.Front Plant Sci. 2020 Jun 30;11:716. doi: 10.3389/fpls.2020.00716. eCollection 2020. Front Plant Sci. 2020. PMID: 32695124 Free PMC article.
-
Predicting Variabilities in Cardiac Gene Expression with a Boolean Network Incorporating Uncertainty.PLoS One. 2015 Jul 24;10(7):e0131832. doi: 10.1371/journal.pone.0131832. eCollection 2015. PLoS One. 2015. PMID: 26207376 Free PMC article.
-
The Wheat GT Factor TaGT2L1D Negatively Regulates Drought Tolerance and Plant Development.Sci Rep. 2016 Jun 1;6:27042. doi: 10.1038/srep27042. Sci Rep. 2016. PMID: 27245096 Free PMC article.
-
Network model of immune responses reveals key effectors to single and co-infection dynamics by a respiratory bacterium and a gastrointestinal helminth.PLoS Comput Biol. 2012 Jan;8(1):e1002345. doi: 10.1371/journal.pcbi.1002345. Epub 2012 Jan 12. PLoS Comput Biol. 2012. PMID: 22253585 Free PMC article.
-
Uncovering signal transduction networks from high-throughput data by integer linear programming.Nucleic Acids Res. 2008 May;36(9):e48. doi: 10.1093/nar/gkn145. Epub 2008 Apr 13. Nucleic Acids Res. 2008. PMID: 18411207 Free PMC article.
References
-
- Fall CP, Marland ES, Wagner JM, Tyson JJ. Computational cell biology. New York: Springer; 2002. 468
-
- Voit EO. Computational analysis of biochemical systems. Cambridge: Cambridge University Press; 2000. 531
-
- Bower JM, Bolouri H. Computational modeling of genetic and biochemical networks. Cambridge (Massachusetts): MIT Press; 2001. 336
-
- Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae . Nature. 2000;403:623–627. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

