Treatment of Chlamydia trachomatis with a small molecule inhibitor of the Yersinia type III secretion system disrupts progression of the chlamydial developmental cycle
- PMID: 16968227
- PMCID: PMC1615999
- DOI: 10.1111/j.1365-2958.2006.05347.x
Treatment of Chlamydia trachomatis with a small molecule inhibitor of the Yersinia type III secretion system disrupts progression of the chlamydial developmental cycle
Abstract
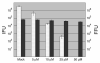
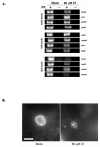
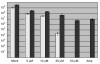
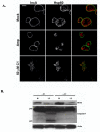
The obligate intracellular bacterium Chlamydia trachomatis possesses a biphasic developmental cycle that is manifested by differentiation of infectious, metabolically inert elementary bodies (EBs) to larger, metabolically active reticulate bodies (RBs). The cycle is completed by asynchronous differentiation of dividing RBs back to a population of dormant EBs that can initiate further rounds of infection upon lysis of the host cell. Chlamydiae express a type III secretion system (T3SS) that is presumably employed to establish and maintain the permissive intracellular niche by secretion of anti-host proteins. We hypothesize that T3SS activity is essential for chlamydial development and pathogenesis. However, the lack of a genetic system has confounded efforts to establish any role of the T3SS. We therefore employed the small molecule Yersinia T3SS inhibitor N'-(3,5-dibromo-2-hydroxybenzylidene)-4-nitrobenzohydrazide, designated compound 1 (C1), to examine the interdependence of the chlamydial T3SS and development. C1 treatment inhibited C. trachomatis but not T4SS-expressing Coxiella burnetii development in a dose-dependent manner. Although chlamydiae remained viable and metabolically active, they failed to divide significantly and RB to EB differentiation was inhibited. These effects occurred in the absence of host cell cytotoxicity and were reversible by washing out C1. We further demonstrate that secretion of T3S substrates is perturbed in C1-treated chlamydial cultures. We have therefore provided evidence that C1 can inhibit C. trachomatis development and T3SS activity and present a model in which progression of the C. trachomatis developmental cycle requires a fully functional T3SS.
Figures


Similar articles
-
Impact of Active Metabolism on Chlamydia trachomatis Elementary Body Transcript Profile and Infectivity.J Bacteriol. 2018 Jun 25;200(14):e00065-18. doi: 10.1128/JB.00065-18. Print 2018 Jul 15. J Bacteriol. 2018. PMID: 29735758 Free PMC article.
-
Type III Secretion in Chlamydia.Microbiol Mol Biol Rev. 2023 Sep 26;87(3):e0003423. doi: 10.1128/mmbr.00034-23. Epub 2023 Jun 26. Microbiol Mol Biol Rev. 2023. PMID: 37358451 Free PMC article. Review.
-
Evidence for the secretion of Chlamydia trachomatis CopN by a type III secretion mechanism.Mol Microbiol. 2000 Dec;38(5):1048-60. doi: 10.1046/j.1365-2958.2000.02212.x. Mol Microbiol. 2000. PMID: 11123678
-
The T3SS structural and effector genes of Chlamydia trachomatis are expressed in distinct phenotypic cell forms.Front Cell Infect Microbiol. 2025 May 8;15:1579247. doi: 10.3389/fcimb.2025.1579247. eCollection 2025. Front Cell Infect Microbiol. 2025. PMID: 40406518 Free PMC article.
-
Host-pathogen reorganisation during host cell entry by Chlamydia trachomatis.Microbes Infect. 2015 Nov-Dec;17(11-12):727-31. doi: 10.1016/j.micinf.2015.08.004. Epub 2015 Aug 28. Microbes Infect. 2015. PMID: 26320027 Free PMC article. Review.
Cited by
-
Salicylidene acylhydrazides that affect type III protein secretion in Salmonella enterica serovar typhimurium.Antimicrob Agents Chemother. 2007 Aug;51(8):2867-76. doi: 10.1128/AAC.00223-07. Epub 2007 Jun 4. Antimicrob Agents Chemother. 2007. PMID: 17548496 Free PMC article.
-
A novel inhibitor of Chlamydophila pneumoniae protein kinase D (PknD) inhibits phosphorylation of CdsD and suppresses bacterial replication.BMC Microbiol. 2009 Oct 14;9:218. doi: 10.1186/1471-2180-9-218. BMC Microbiol. 2009. PMID: 19828035 Free PMC article.
-
Synthesis of [4-(2-hydroxyphenyl)thiazol-2-yl]methanones as potential bioisosteres of salicylidene acylhydrazides.Molecules. 2010 Aug 31;15(9):6019-34. doi: 10.3390/molecules15096019. Molecules. 2010. PMID: 20877207 Free PMC article.
-
From screen to target: insights and approaches for the development of anti-virulence compounds.Front Cell Infect Microbiol. 2014 Sep 30;4:139. doi: 10.3389/fcimb.2014.00139. eCollection 2014. Front Cell Infect Microbiol. 2014. PMID: 25325019 Free PMC article. Review.
-
Mutations in hemG mediate resistance to salicylidene acylhydrazides, demonstrating a novel link between protoporphyrinogen oxidase (HemG) and Chlamydia trachomatis infectivity.J Bacteriol. 2013 Sep;195(18):4221-30. doi: 10.1128/JB.00506-13. Epub 2013 Jul 12. J Bacteriol. 2013. PMID: 23852872 Free PMC article.
References
-
- Clausen JD, Christiansen G, Holst HU, Birkelund S. Chlamydia trachomatis utilizes the host cell microtubule network during early events of infection. Mol Microbiol. 1997;25:441–449. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

