An antibody binding site on cytochrome c defined by hydrogen exchange and two-dimensional NMR
- PMID: 1697101
- PMCID: PMC3432411
- DOI: 10.1126/science.1697101
An antibody binding site on cytochrome c defined by hydrogen exchange and two-dimensional NMR
Abstract
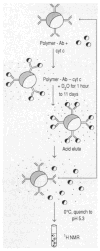
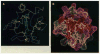
The interaction of a protein antigen, horse cytochrome c (cyt c), with a monoclonal antibody has been studied by hydrogen-deuterium (H-D) exchange labeling and two-dimensional nuclear magnetic resonance (2D NMR) methods. The H-exchange rate of residues in three discontiguous regions of the cyt c polypeptide backbone was slowed by factors up to 340-fold in the antibody-antigen complex compared with free cyt c. The protected residues, 36 to 38, 59, 60, 64 to 67, 100, and 101, and their hydrogen-bond acceptors, are brought together in the three-dimensional structure to form a contiguous, largely exposed protein surface with an area of about 750 square angstroms. The interaction site determined in this way is consistent with prior epitope mapping studies and includes several residues that were not previously identified. The hydrogen exchange labeling approach can be used to map binding sites on small proteins in antibody-antigen complexes and may be applicable to protein-protein and protein-ligand interactions in general.
Figures


References
-
- Davies DR, Sheriff S, Padlan EA. J Biol Chem. 1988;263:10,541. - PubMed
-
- Benjamin DC, et al. Annu Rev Immunol. 1984;2:67. - PubMed
- Jemmerson R, Paterson Y. Bio Techniques. 1986;4:1818.
-
- Cooper HM, et al. J Biol Chem. 1987;262:11,591.
-
- Collawn JF, Wallace CJA, Proudfoot AEI, Paterson Y. 1988;263:8625. ibid. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

