One protein, two enzymes revisited: a structural entropy switch interconverts the two isoforms of acireductone dioxygenase
- PMID: 16989860
- PMCID: PMC1808343
- DOI: 10.1016/j.jmb.2006.08.060
One protein, two enzymes revisited: a structural entropy switch interconverts the two isoforms of acireductone dioxygenase
Abstract
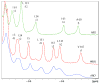
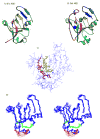
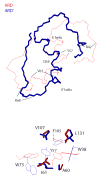
Acireductone dioxygenase (ARD) catalyzes different reactions between O2 and 1,2-dihydroxy-3-oxo-5-(methylthio)pent-1-ene (acireductone) depending upon the metal bound in the active site. Ni2+ -ARD cleaves acireductone to formate, CO and methylthiopropionate. If Fe2+ is bound (ARD'), the same substrates yield methylthioketobutyrate and formate. The two forms differ in structure, and are chromatographically separable. Paramagnetism of Fe2+ renders the active site of ARD' inaccessible to standard NMR methods. The structure of ARD' has been determined using Fe2+ binding parameters determined by X-ray absorption spectroscopy and NMR restraints from H98S ARD, a metal-free diamagnetic protein that is isostructural with ARD'. ARD' retains the beta-sandwich fold of ARD, but a structural entropy switch increases order at one end of a two-helix system that bisects the beta-sandwich and decreases order at the other upon interconversion of ARD and ARD', causing loss of the C-terminal helix in ARD' and rearrangements of residues involved in substrate orientation in the active site.
Figures

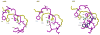


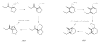
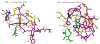
Similar articles
-
Characterization of metal binding in the active sites of acireductone dioxygenase isoforms from Klebsiella ATCC 8724.Biochemistry. 2008 Feb 26;47(8):2428-38. doi: 10.1021/bi7004152. Epub 2008 Feb 1. Biochemistry. 2008. PMID: 18237192 Free PMC article.
-
Metal-dependent activity of Fe and Ni acireductone dioxygenases: how two electrons reroute the catalytic pathway.J Mol Biol. 2013 Aug 23;425(16):3007-18. doi: 10.1016/j.jmb.2013.05.001. Epub 2013 May 13. J Mol Biol. 2013. PMID: 23680285
-
Dual chemistry catalyzed by human acireductone dioxygenase.Protein Eng Des Sel. 2017 Mar 1;30(3):197-204. doi: 10.1093/protein/gzw078. Protein Eng Des Sel. 2017. PMID: 28062648 Free PMC article.
-
The Metal Drives the Chemistry: Dual Functions of Acireductone Dioxygenase.Chem Rev. 2017 Aug 9;117(15):10474-10501. doi: 10.1021/acs.chemrev.7b00117. Epub 2017 Jul 21. Chem Rev. 2017. PMID: 28731690 Free PMC article. Review.
-
Non-heme iron-dependent dioxygenases: unravelling catalytic mechanisms for complex enzymatic oxidations.Curr Opin Chem Biol. 2008 Apr;12(2):134-40. doi: 10.1016/j.cbpa.2007.12.007. Epub 2008 Feb 20. Curr Opin Chem Biol. 2008. PMID: 18249197 Review.
Cited by
-
Probing the mechanism of recognition of ssDNA by the Cdc13-DBD.Nucleic Acids Res. 2008 Mar;36(5):1624-33. doi: 10.1093/nar/gkn017. Epub 2008 Feb 3. Nucleic Acids Res. 2008. PMID: 18250086 Free PMC article.
-
Characterization of metal binding in the active sites of acireductone dioxygenase isoforms from Klebsiella ATCC 8724.Biochemistry. 2008 Feb 26;47(8):2428-38. doi: 10.1021/bi7004152. Epub 2008 Feb 1. Biochemistry. 2008. PMID: 18237192 Free PMC article.
-
O2-dependent aliphatic carbon-carbon bond cleavage reactivity in a Ni(II) enolate complex having a hydrogen bond donor microenvironment; comparison with a hydrophobic analogue.Inorg Chem. 2010 Feb 1;49(3):1071-81. doi: 10.1021/ic901981y. Inorg Chem. 2010. PMID: 20039645 Free PMC article.
-
Nickel ions inhibit histone demethylase JMJD1A and DNA repair enzyme ABH2 by replacing the ferrous iron in the catalytic centers.J Biol Chem. 2010 Mar 5;285(10):7374-83. doi: 10.1074/jbc.M109.058503. Epub 2009 Dec 30. J Biol Chem. 2010. PMID: 20042601 Free PMC article.
-
Ring-cleaving dioxygenases with a cupin fold.Appl Environ Microbiol. 2012 Apr;78(8):2505-14. doi: 10.1128/AEM.07651-11. Epub 2012 Jan 27. Appl Environ Microbiol. 2012. PMID: 22287012 Free PMC article. Review.
References
-
- deAraujo AFP, Pochapsky TC. Monte Carlo simulations of protein folding using inexact potentials: How accurate must parameters be in order to preserve the essential features of the energy landscape? Folding & Design. 1996;1:299–314. - PubMed
-
- Shakhnovich EI. Proteins with selected sequences fold into unique native conformation. Phys Rev Lett. 1994;72:3907–3910. - PubMed
-
- Devlin GL, Bottomley SP. A protein family under ‘stress’ - Serpin stability, folding and misfolding. Frontiers In Bioscience. 2005;10:288–299. - PubMed
-
- Wray JW, Abeles RH. A bacterial enzyme that catalyzes formation of carbon monoxide. J Biol Chem. 1993;268:21466–21469. - PubMed
References
-
- Dai Y, Wensink PC, Abeles RH. One protein, two enzymes. Journal Of Biological Chemistry. 1999;274:1193–1195. - PubMed
-
- Roe AL, Schneider DJ, Mayer RJ, Pyrz JW, Widom J, Que L. X-Ray Absorption-Spectroscopy of Iron-Tyrosinate Proteins. Journal of the American Chemical Society. 1984;106:1676–1681.
-
- Randall CR, Zang Y, True AE, Que L, Charnock JM, Garner CD, Fujishima Y, Schofield CJ, Baldwin JE. X-Ray-Absorption Studies of the Ferrous Active-Site of Isopenicillin N-Synthase and Related Model Complexes. Biochemistry. 1993;32:6664–6673. - PubMed
-
- Bertini I, Briganti F, Mangani S, Nolting HF, Scozzafava A. X-Ray-Absorption Studies on Catechol 2,3-Dioxygenase from Pseudomonas-Putida Mt2. Biochemistry. 1994;33:10777–10784. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

