PDV1 and PDV2 mediate recruitment of the dynamin-related protein ARC5 to the plastid division site
- PMID: 16998069
- PMCID: PMC1626610
- DOI: 10.1105/tpc.106.045484
PDV1 and PDV2 mediate recruitment of the dynamin-related protein ARC5 to the plastid division site
Abstract
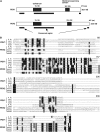
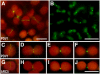
During plastid division, the dynamin-related protein ACCUMULATION AND REPLICATION OF CHLOROPLASTS5 (ARC5) is recruited from the cytosol to the surface of the outer chloroplast envelope membrane. In Arabidopsis thaliana arc5 mutants, chloroplasts arrest during division site constriction. Analysis of mutants similar to arc5 along with map-based cloning identified PLASTID DIVISION1 (PDV1), an integral outer envelope membrane protein, and its homolog PDV2 as components of the plastid division machinery. Similar to ARC5, PDV1 localized to a discontinuous ring at the division site in wild-type plants. The midplastid PDV1 ring formed in arc5 mutants and the ARC5 ring formed in pdv1 and pdv2 mutants, but not in pdv1 pdv2. Stromal FtsZ ring assembly occurred in pdv1, pdv2, and pdv1 pdv2, as it does in arc5. Topological analysis showed that the large N-terminal region of PDV1 upstream of the transmembrane helix bearing a putative coiled-coil domain is exposed to the cytosol. Mutation of the conserved PDV1 C-terminal Gly residue did not block PDV1 insertion into the outer envelope membrane but did abolish its localization to the division site. Our results indicate that plastid division involves the stepwise localization of FtsZ, PDV1, and ARC5 at the division site and that PDV1 and PDV2 together mediate the recruitment of ARC5 to the midplastid constriction at a late stage of division.
Figures







Comment in
-
Pushing the envelope: the role of outer envelope proteins PVD1 and PVD2 in plastid division.Plant Cell. 2006 Oct;18(10):2419. doi: 10.1105/tpc.106.181011. Plant Cell. 2006. PMID: 17088605 Free PMC article. No abstract available.
References
-
- Aldridge, C., Maple, J., and Møller, S.G. (2005). The molecular biology of plastid division in higher plants. J. Exp. Bot. 56 1061–1077. - PubMed
-
- Asano, T., Yoshioka, Y., Kurei, S., Sakamoto, W., Sodmergen, and Machida, Y. (2004). A mutation of the CRUMPLED LEAF gene that encodes a protein localized in the outer envelope membrane of plastids affects the pattern of cell division, cell differentiation, and plastid division in Arabidopsis. Plant J. 38 448–459. - PubMed
-
- Bell, C.J., and Ecker, J.R. (1994). Assignment of 30 microsatellite loci to the linkage map of Arabidopsis. Genomics 19 137–144. - PubMed
-
- Bhar, D., Karren, M.A., Babst, M., and Shaw, J.M. (2006). Dimeric Dnm1–G385D interacts with Mdv1 on mitochondria and can be stimulated to assemble into fission complexes containing Mdv1 and Fis1. J. Biol. Chem. 281 17312–17320. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

