Sequence identity of the direct repeats, DR1 and DR2, contributes to the discrimination between primer translocation and in situ priming during replication of the duck hepatitis B virus
- PMID: 17005197
- PMCID: PMC1803024
- DOI: 10.1016/j.jmb.2006.08.095
Sequence identity of the direct repeats, DR1 and DR2, contributes to the discrimination between primer translocation and in situ priming during replication of the duck hepatitis B virus
Abstract
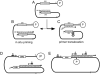
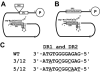
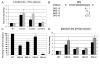
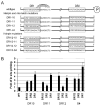
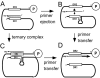
There are two mutually exclusive pathways for plus-strand DNA synthesis in hepadnavirus reverse transcription. The predominant pathway gives rise to relaxed circular DNA, while the other pathway yields duplex linear DNA. At the completion of minus-strand DNA synthesis, the final RNase H cleavage generates the plus-strand primer at direct repeat 1 (DR1). A small fraction of viruses make duplex linear DNA after initiating plus-strand DNA synthesis from this site, a process called in situ priming. To make relaxed circular DNA, a template switch is necessary for the RNA primer generated at DR1 to initiate plus-strand DNA synthesis from the direct repeat 2 (DR2) located near the opposite end of the minus-strand DNA, a process called primer translocation. We are interested in understanding the mechanism that discriminates between these two processes. Previously, we showed that a small DNA hairpin forms at DR1 in the avihepadnaviruses and acts as an inhibitor of in situ priming. Here, using genetic approaches, we show that sequence identity between DR1 and DR2 is necessary, but not sufficient for primer translocation in the duck hepatitis B virus. The discrimination between in situ priming and primer translocation depends upon suppression of in situ priming, a process that is dependent upon both sequence identity between DR1 and DR2, and the presence of the hairpin at DR1. Finally, our analysis indicates the entire RNA primer can contribute to primer translocation and is translocated to DR2 before initiation of plus-strand DNA synthesis from that site.
Figures


Similar articles
-
Small DNA hairpin negatively regulates in situ priming during duck hepatitis B virus reverse transcription.J Virol. 2002 Feb;76(3):980-9. doi: 10.1128/jvi.76.3.980-989.2002. J Virol. 2002. PMID: 11773373 Free PMC article.
-
Mutations affecting hepadnavirus plus-strand DNA synthesis dissociate primer cleavage from translocation and reveal the origin of linear viral DNA.J Virol. 1991 Mar;65(3):1255-62. doi: 10.1128/JVI.65.3.1255-1262.1991. J Virol. 1991. PMID: 1704925 Free PMC article.
-
Mutations within DR2 independently reduce the amount of both minus- and plus-strand DNA synthesized during duck hepatitis B virus replication.J Virol. 1996 Dec;70(12):8684-90. doi: 10.1128/JVI.70.12.8684-8690.1996. J Virol. 1996. PMID: 8970995 Free PMC article.
-
Analysis of duck hepatitis B virus reverse transcription indicates a common mechanism for the two template switches during plus-strand DNA synthesis.J Virol. 2002 Mar;76(6):2763-9. doi: 10.1128/jvi.76.6.2763-2769.2002. J Virol. 2002. PMID: 11861843 Free PMC article.
-
Hepatitis B virus replication.World J Gastroenterol. 2007 Jan 7;13(1):48-64. doi: 10.3748/wjg.v13.i1.48. World J Gastroenterol. 2007. PMID: 17206754 Free PMC article. Review.
Cited by
-
The interface between hepatitis B virus capsid proteins affects self-assembly, pregenomic RNA packaging, and reverse transcription.J Virol. 2015 Mar;89(6):3275-84. doi: 10.1128/JVI.03545-14. Epub 2015 Jan 7. J Virol. 2015. PMID: 25568211 Free PMC article.
-
The sequence of the RNA primer and the DNA template influence the initiation of plus-strand DNA synthesis in hepatitis B virus.J Mol Biol. 2007 Jul 13;370(3):471-80. doi: 10.1016/j.jmb.2007.04.057. Epub 2007 May 4. J Mol Biol. 2007. PMID: 17531265 Free PMC article.
-
Human hepatitis B virus production in avian cells is characterized by enhanced RNA splicing and the presence of capsids containing shortened genomes.PLoS One. 2012;7(5):e37248. doi: 10.1371/journal.pone.0037248. Epub 2012 May 18. PLoS One. 2012. PMID: 22624002 Free PMC article.
-
Insights into Immune Exhaustion in Chronic Hepatitis B: A Review of Checkpoint Receptor Expression.Pharmaceuticals (Basel). 2024 Jul 21;17(7):964. doi: 10.3390/ph17070964. Pharmaceuticals (Basel). 2024. PMID: 39065812 Free PMC article. Review.
References
-
- Goff SP. Retroviridae: The Retroviruses and Their Replication. In: Knipe DM, H PM, et al., editors. Fields Virology. 4. Vol. 2. Lippincott-Raven; Philadelphia, PA: 2001. pp. 1871–1939. 2 vols.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

