Effects of picornavirus 3A Proteins on Protein Transport and GBF1-dependent COP-I recruitment
- PMID: 17005635
- PMCID: PMC1642585
- DOI: 10.1128/JVI.01225-06
Effects of picornavirus 3A Proteins on Protein Transport and GBF1-dependent COP-I recruitment
Abstract
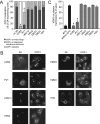
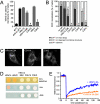
The 3A protein of the coxsackievirus B3 (CVB3), an enterovirus that belongs to the family of the picornaviruses, inhibits endoplasmic reticulum-to-Golgi transport. Recently, we elucidated the underlying mechanism by showing that CVB3 3A interferes with ADP-ribosylation factor 1 (Arf1)-dependent COP-I recruitment to membranes by binding and inhibiting the function of GBF1, a guanine nucleotide exchange factor that is required for the activation of Arf1 (E. Wessels et al., Dev. Cell 11:191-201, 2006). Here, we show that the 3A protein of poliovirus, another enterovirus, is also able to interfere with COP-I recruitment through the same mechanism. No interference with protein transport or COP-I recruitment was observed for the 3A proteins of any of the other picornaviruses tested here (human rhinovirus [HRV], encephalomyocarditis virus, foot-and-mouth disease virus, and hepatitis A virus). We show that the 3A proteins of HRV, which are the most closely related to the enteroviruses, are unable to inhibit COP-I recruitment, due to a reduced ability to bind GBF1. When the N-terminal residues of the HRV 3A proteins are replaced by those of CVB3 3A, chimeric proteins are produced that have gained the ability to bind GBF1 and, by consequence, to inhibit protein transport. These results show that the N terminus of the CVB3 3A protein is important for binding of GBF1 and its transport-inhibiting function. Taken together, our data demonstrate that the activity of the enterovirus 3A protein to inhibit GBF1-dependent COP-I recruitment is unique among the picornaviruses.
Figures

References
-
- Bienz, K., D. Egger, and T. Pfister. 1994. Characteristics of the poliovirus replication complex. Arch. Virol. 9(Suppl.):147-157. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

