Characterization of a novel 5' subgenomic RNA3a derived from RNA3 of Brome mosaic bromovirus
- PMID: 17005659
- PMCID: PMC1676258
- DOI: 10.1128/JVI.01207-06
Characterization of a novel 5' subgenomic RNA3a derived from RNA3 of Brome mosaic bromovirus
Abstract
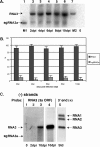
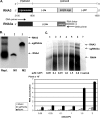
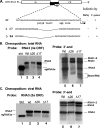
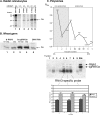
The synthesis of 3' subgenomic RNA4 (sgRNA4) by initiation from an internal sg promoter in the RNA3 segment was first described for Brome mosaic bromovirus (BMV), a model tripartite positive-sense RNA virus (W. A. Miller, T. W. Dreher, and T. C. Hall, Nature 313:68-70, 1985). In this work, we describe a novel 5' sgRNA of BMV (sgRNA3a) that we propose arises by premature internal termination and that encapsidates in BMV virions. Cloning and sequencing revealed that, unlike any other BMV RNA segment, sgRNA3a carries a 3' oligo(A) tail, in which respect it resembles cellular mRNAs. Indeed, both the accumulation of sgRNA3a in polysomes and the synthesis of movement protein 3a in in vitro systems suggest active functions of sgRNA3a during protein synthesis. Moreover, when copied in the BMV replicase in vitro reaction, the minus-strand RNA3 template generated the sgRNA3a product, likely by premature termination at the minus-strand oligo(U) tract. Deletion of the oligo(A) tract in BMV RNA3 inhibited synthesis of sgRNA3a during infection. We propose a model in which the synthesis of RNA3 is terminated prematurely near the sg promoter. The discovery of 5' sgRNA3a sheds new light on strategies viruses can use to separate replication from the translation functions of their genomic RNAs.
Figures


Similar articles
-
Recombination of 5' subgenomic RNA3a with genomic RNA3 of Brome mosaic bromovirus in vitro and in vivo.Virology. 2011 Feb 5;410(1):129-41. doi: 10.1016/j.virol.2010.10.037. Epub 2010 Nov 26. Virology. 2011. PMID: 21111438 Free PMC article.
-
Second-site mutations in the brome mosaic virus RNA3 intercistronic region partially suppress a defect in coat protein mRNA transcription.Virology. 1994 Feb;198(2):427-36. doi: 10.1006/viro.1994.1054. Virology. 1994. PMID: 8291227
-
Replicase-binding sites on plus- and minus-strand brome mosaic virus RNAs and their roles in RNA replication in plant cells.J Virol. 2004 Dec;78(24):13420-9. doi: 10.1128/JVI.78.24.13420-13429.2004. J Virol. 2004. PMID: 15564452 Free PMC article.
-
The brome mosaic virus 3' untranslated sequence regulates RNA replication, recombination, and virion assembly.Virus Res. 2015 Aug 3;206:46-52. doi: 10.1016/j.virusres.2015.02.007. Epub 2015 Feb 14. Virus Res. 2015. PMID: 25687214 Review.
-
Common replication strategies emerging from the study of diverse groups of positive-strand RNA viruses.Arch Virol Suppl. 1994;9:181-94. doi: 10.1007/978-3-7091-9326-6_18. Arch Virol Suppl. 1994. PMID: 8032249 Review.
Cited by
-
Subgenomic messenger RNAs: mastering regulation of (+)-strand RNA virus life cycle.Virology. 2011 Apr 10;412(2):245-55. doi: 10.1016/j.virol.2011.02.007. Epub 2011 Mar 5. Virology. 2011. PMID: 21377709 Free PMC article. Review.
-
Insights into the single-cell reproduction cycle of members of the family Bromoviridae: lessons from the use of protoplast systems.J Virol. 2008 Nov;82(21):10330-40. doi: 10.1128/JVI.00746-08. Epub 2008 Aug 6. J Virol. 2008. PMID: 18684833 Free PMC article. Review. No abstract available.
-
Long Noncoding RNAs in Plant Viroids and Viruses: A Review.Pathogens. 2020 Sep 18;9(9):765. doi: 10.3390/pathogens9090765. Pathogens. 2020. PMID: 32961969 Free PMC article. Review.
-
The plant host can affect the encapsidation of brome mosaic virus (BMV) RNA: BMV virions are surprisingly heterogeneous.J Mol Biol. 2014 Mar 6;426(5):1061-76. doi: 10.1016/j.jmb.2013.09.007. Epub 2013 Sep 13. J Mol Biol. 2014. PMID: 24036424 Free PMC article.
-
Brome mosaic virus capsid protein regulates accumulation of viral replication proteins by binding to the replicase assembly RNA element.RNA. 2009 Apr;15(4):615-26. doi: 10.1261/rna.1375509. Epub 2009 Feb 23. RNA. 2009. PMID: 19237464 Free PMC article.
References
-
- Ahlquist, P., R. Dasgupta, and P. Kaesberg. 1984. Nucleotide sequence of the Brome mosaic virus genome and its implications for viral replication. J. Mol. Biol. 172:369-383. - PubMed
-
- Ayllon, M. A., S. Gowda, T. Satyanarayana, and W. O. Dawson. 2004. cis-acting elements at opposite ends of the Citrus tristeza virus genome differ in initiation and termination of subgenomic RNAs. Virology 322:41-50. - PubMed
-
- Beelman, C., and R. Parker. 1995. Degradation of mRNA in eukaryotes. Cell 81:179-183. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

