Substrate recognition, protein dynamics, and iron-sulfur cluster in Pseudomonas aeruginosa adenosine 5'-phosphosulfate reductase
- PMID: 17010373
- PMCID: PMC1769331
- DOI: 10.1016/j.jmb.2006.08.080
Substrate recognition, protein dynamics, and iron-sulfur cluster in Pseudomonas aeruginosa adenosine 5'-phosphosulfate reductase
Abstract
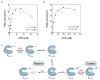
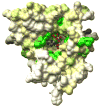
APS reductase catalyzes the first committed step of reductive sulfate assimilation in pathogenic bacteria, including Mycobacterium tuberculosis, and is a promising target for drug development. We report the 2.7 A resolution crystal structure of Pseudomonas aeruginosa APS reductase in the thiosulfonate intermediate form of the catalytic cycle and with substrate bound. The structure, high-resolution Fourier transform ion cyclotron resonance (FT-ICR) mass spectrometry, and quantitative kinetic analysis, establish that the two chemically discrete steps of the overall reaction take place at distinct sites on the enzyme, mediated via conformational flexibility of the C-terminal 18 residues. The results address the mechanism by which sulfonucleotide reductases protect the covalent but labile enzyme-intermediate before release of sulfite by the protein cofactor thioredoxin. P. aeruginosa APS reductase contains an [4Fe-4S] cluster that is essential for catalysis. The structure reveals an unusual mode of cluster coordination by tandem cysteine residues and suggests how this arrangement might facilitate conformational change and cluster interaction with the substrate. Assimilatory 3'-phosphoadenosine 5'-phosphosulfate (PAPS) reductases are evolutionarily related, homologous enzymes that catalyze the same overall reaction, but do so in the absence of an [Fe-S] cluster. The APS reductase structure reveals adaptive use of a phosphate-binding loop for recognition of the APS O3' hydroxyl group, or the PAPS 3'-phosphate group.
Figures
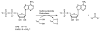
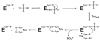

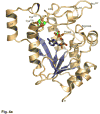
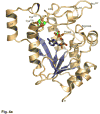
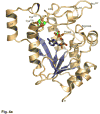







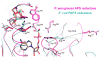
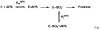
References
-
- Kopriva S, Buchert T, Fritz G, Suter M, Weber M, Benda R, Schaller J, Feller U, Schurmann P, Schunemann V, Trautwein AX, Kroneck PM, Brunold C. Plant adenosine 5′-phosphosulfate reductase is a novel iron-sulfur protein. J Biol Chem. 2001;276:42881–42886. - PubMed
-
- Williams SJ, Senaratne RH, Mougous JD, Riley LW, Bertozzi CR. 5′-Adenosinephosphosulfate lies at a metabolic branch point in mycobacteria. J Biol Chem. 2002;277:32606–32615. - PubMed
-
- Leyh TS, Taylor JC, Markham GD. The sulfate activation locus of Escherichia coli K12: cloning, genetic, and enzymatic characterization. J Biol Chem. 1988;263:2409–2416. - PubMed
-
- Mougous JD, Lee DH, Hubbard SC, Schelle MW, Vocadlo DJ, Berger JM, Bertozzi CR. Molecular basis for G protein control of the prokaryotic ATP sulfurylase. Mol Cell. 2006;21:109–122. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

