SarA is a repressor of hla (alpha-hemolysin) transcription in Staphylococcus aureus: its apparent role as an activator of hla in the prototype strain NCTC 8325 depends on reduced expression of sarS
- PMID: 17012389
- PMCID: PMC1698246
- DOI: 10.1128/JB.00866-06
SarA is a repressor of hla (alpha-hemolysin) transcription in Staphylococcus aureus: its apparent role as an activator of hla in the prototype strain NCTC 8325 depends on reduced expression of sarS
Abstract
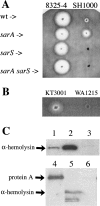
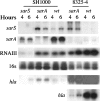
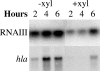
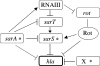
In most Staphylococcus aureus strains, inactivation of sarA increases hla transcription, indicating that sarA is a repressor. However, in S. aureus NCTC 8325 and its derivatives, used for most studies of hla regulation, inactivation of sarA resulted in decreased hla transcription. The disparate phenotype of strain NCTC 8325 seems to be associated with its rsbU mutation, which leads to sigma(B) deficiency. This has now been verified by the demonstration that sarA repressed hla transcription in an rsbU+ derivative of strain 8325-4 (SH1000). That sarA could act as a repressor of hla in an 8325-4 background was confirmed by the observation that inactivation of sarA in an agr sarS rot triple mutant dramatically increased hla transcription to wild-type levels. However, the apparent role of sarA as an activator of hla in 8325-4 was not a result of the rsbU mutation alone, as inactivation of sarA in another rsbU mutant, strain V8, led to increased hla transcription. Northern blot analysis revealed much higher levels of sarS mRNA in strain V8 than in 8325-4, which was likely due to the mutation in the sarS activator, tcaR, in 8325-4, which was not found in strain V8. On the other hand, the relative increase in sarS transcription upon the inactivation of sarA was 15-fold higher in 8325-4 than in strain V8. Because of this, inactivation of sarA in 8325-4 means a net increase in repressor activity, whereas in strain V8, inactivation of sarA means a net decrease in repressor activity and, therefore, enhanced hla transcription.
Figures





Similar articles
-
Defining the strain-dependent impact of the Staphylococcal accessory regulator (sarA) on the alpha-toxin phenotype of Staphylococcus aureus.J Bacteriol. 2011 Jun;193(12):2948-58. doi: 10.1128/JB.01517-10. Epub 2011 Apr 8. J Bacteriol. 2011. PMID: 21478342 Free PMC article.
-
Rat/MgrA, a regulator of autolysis, is a regulator of virulence genes in Staphylococcus aureus.Infect Immun. 2005 Mar;73(3):1423-31. doi: 10.1128/IAI.73.3.1423-1431.2005. Infect Immun. 2005. PMID: 15731040 Free PMC article.
-
SarT, a repressor of alpha-hemolysin in Staphylococcus aureus.Infect Immun. 2001 Aug;69(8):4749-58. doi: 10.1128/IAI.69.8.4749-4758.2001. Infect Immun. 2001. PMID: 11447147 Free PMC article.
-
sigmaB modulates virulence determinant expression and stress resistance: characterization of a functional rsbU strain derived from Staphylococcus aureus 8325-4.J Bacteriol. 2002 Oct;184(19):5457-67. doi: 10.1128/JB.184.19.5457-5467.2002. J Bacteriol. 2002. PMID: 12218034 Free PMC article.
-
The SarA protein family of Staphylococcus aureus.Int J Biochem Cell Biol. 2008;40(3):355-61. doi: 10.1016/j.biocel.2007.10.032. Epub 2007 Nov 13. Int J Biochem Cell Biol. 2008. PMID: 18083623 Free PMC article. Review.
Cited by
-
Eugenol reduces the expression of virulence-related exoproteins in Staphylococcus aureus.Appl Environ Microbiol. 2010 Sep;76(17):5846-51. doi: 10.1128/AEM.00704-10. Epub 2010 Jul 16. Appl Environ Microbiol. 2010. PMID: 20639367 Free PMC article.
-
Identification of single nucleotide polymorphisms associated with hyperproduction of alpha-toxin in Staphylococcus aureus.PLoS One. 2011 Apr 8;6(4):e18428. doi: 10.1371/journal.pone.0018428. PLoS One. 2011. PMID: 21494631 Free PMC article.
-
Impact of the functional status of saeRS on in vivo phenotypes of Staphylococcus aureus sarA mutants.Mol Microbiol. 2014 Jun;92(6):1299-312. doi: 10.1111/mmi.12629. Epub 2014 May 12. Mol Microbiol. 2014. PMID: 24779437 Free PMC article.
-
Defining the strain-dependent impact of the Staphylococcal accessory regulator (sarA) on the alpha-toxin phenotype of Staphylococcus aureus.J Bacteriol. 2011 Jun;193(12):2948-58. doi: 10.1128/JB.01517-10. Epub 2011 Apr 8. J Bacteriol. 2011. PMID: 21478342 Free PMC article.
-
Epistatic relationships between sarA and agr in Staphylococcus aureus biofilm formation.PLoS One. 2010 May 24;5(5):e10790. doi: 10.1371/journal.pone.0010790. PLoS One. 2010. PMID: 20520723 Free PMC article.
References
-
- Arvidson, S. 2004. Virulence gene regulation and their role in pathogenesis of disease, p. 154-176. In D. Ala'Aldeen and K. Hiramatsu (ed.), Staphylococcus aureus, molecular and clinical aspects. Horwood Publishing, Chichester, United Kingdom.
-
- Arvidson, S., and K. Tegmark. 2001. Regulation of virulence determinants in Staphylococcus aureus. Int. J. Med. Microbiol. 291:159-170. - PubMed
-
- Björklind, A., and S. Arvidson. 1980. Mutants of Staphylococcus aureus affected in the regulation of exoprotein synthesis. FEMS Microbiol. Lett. 7:202-206.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

