Intrinsic and selected resistance to antibiotics binding the ribosome: analyses of Brucella 23S rrn, L4, L22, EF-Tu1, EF-Tu2, efflux and phylogenetic implications
- PMID: 17014718
- PMCID: PMC1617103
- DOI: 10.1186/1471-2180-6-84
Intrinsic and selected resistance to antibiotics binding the ribosome: analyses of Brucella 23S rrn, L4, L22, EF-Tu1, EF-Tu2, efflux and phylogenetic implications
Abstract
Background: Brucella spp. are highly similar, having identical 16S RNA. However, they have important phenotypic differences such as differential susceptibility to antibiotics binding the ribosome. Neither the differential susceptibility nor its basis has been rigorously studied. Differences found among other conserved ribosomal loci could further define the relationships among the classical Brucella spp.
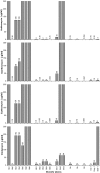
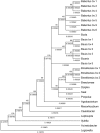
Results: Minimum inhibitory concentration (MIC) values of Brucella reference strains and three marine isolates to antibiotics binding the ribosome ranged from 0.032 to >256 microg/ml for the macrolides erythromycin, clarithromycin, and azithromycin and 2 to >256 microg/ml for the lincosamide, clindamycin. Though sequence polymorphisms were identified among ribosome associated loci 23S rrn, rplV, tuf-1 and tuf-2 but not rplD, they did not correlate with antibiotic resistance phenotypes. When spontaneous erythromycin resistant (eryR) mutants were examined, mutation of the peptidyl transferase center (A2058G Ec) correlated with increased resistance to both erythromycin and clindamycin. Brucella efflux was examined as an alternative antibiotic resistance mechanism by use of the inhibitor L-phenylalanine-L-arginine beta-naphthylamide (PAbetaN). Erythromycin MIC values of reference and all eryR strains, except the B. suis eryR mutants, were lowered variably by PAbetaN. A phylogenetic tree based on concatenated ribosomal associated loci supported separate evolutionary paths for B. abortus, B. melitensis, and B. suis/B. canis, clustering marine Brucella and B. neotomae with B. melitensis. Though Brucella ovis was clustered with B. abortus, the bootstrap value was low.
Conclusion: Polymorphisms among ribosomal loci from the reference Brucella do not correlate with their highly differential susceptibility to erythromycin. Efflux plays an important role in Brucella sensitivity to erythromycin. Polymorphisms identified among ribosome associated loci construct a robust phylogenetic tree supporting classical Brucella spp. designations.
Figures

References
-
- Meyer ME. The epizootiology of brucellosis and its relationship to the identification of Brucella organisms. Am J Vet Res. 1964;25:553–557. - PubMed
-
- Meyer ME. Host-parasite relationships in brucellosis I. reservoirs of infection and interhost transmissibility of the parasite. 70th Proc Annu Meet U S Livestock Sanit Assoc. 1966. pp. 129–136.
-
- Alton GG, Jones LM, Angus RD, Verger JM. Techniques for the Brucellosis Laboratory. Nouzilly, France , Institu National de la Recherche; 1988.
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

