A eukaryotic factor required for accumulation of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis
- PMID: 17041026
- PMCID: PMC1676045
- DOI: 10.1104/pp.106.088682
A eukaryotic factor required for accumulation of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis
Abstract
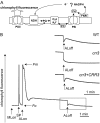
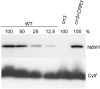
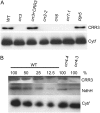
The NAD(P)H dehydrogenase (NDH) complex in chloroplasts mediates photosystem I cyclic and chlororespiratory electron transport. Eleven chloroplast genes and three nuclear genes have been identified as encoding Ndh subunits, but the entire subunit composition is still unknown. An Arabidopsis (Arabidopsis thaliana) chlororespiratory reduction (crr3) mutant was isolated based on its lack of transient increase in chlorophyll fluorescence after actinic light illumination; this was due to a specific defect in accumulation of the NDH complex. The CRR3 gene (At2g01590) encodes a novel protein containing a putative plastid-targeting signal and a transmembrane domain. Consistent with the gene structure, CRR3 localized to the membrane fraction of chloroplasts. In addition to the essential function of CRR3 in stabilizing the NDH complex, the NDH complex is also required for the accumulation of CRR3. These results suggest that CRR3 interacts with the NDH complex in the thylakoid membrane. In contrast to other subunits in the chloroplast NDH complex, CRR3 is not conserved in cyanobacteria from which the chloroplast NDH complex is believed to have originated. We propose that CRR3 is a subunit of the NDH complex, which is specific to the chloroplast.
Figures
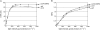


References
-
- Battchikova N, Zhang P, Rudd S, Ogawa T, Aro E-M (2005) Identification of NdhL and Ssl1690 (NdhO) in NDH-1L and NDH-1M complexes of Synechocystis sp. PCC 6803. J Biol Chem 280: 2587–2595 - PubMed
-
- Casano LM, Zapata JM, Martin M, Sabater B (2000) Chlororespiration and poising of cyclic electron transport: plastoquinone as electron transporter between thylakoid NADH dehydrogenase and peroxidase. J Biol Chem 275: 942–948 - PubMed
-
- Choquet Y, Vallon O (2000) Synthesis, assembly and degradation of thylakoid membrane proteins. Biochimie 82: 615–634 - PubMed
-
- De Las Rivas J, Balsera M, Barber J (2004) Evolution of oxygenic photosynthesis: genome-wide analysis of the OEC extrinsic proteins. Trends Plant Sci 19: 18–25 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

