A feedback transcriptional mechanism controls the level of the arginine/lysine transporter cat-1 during amino acid starvation
- PMID: 17042743
- PMCID: PMC1783987
- DOI: 10.1042/BJ20060941
A feedback transcriptional mechanism controls the level of the arginine/lysine transporter cat-1 during amino acid starvation
Abstract
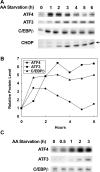
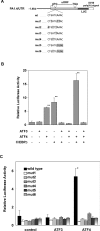
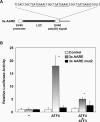
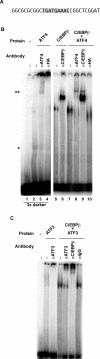
The adaptive response to amino acid limitation in mammalian cells inhibits global protein synthesis and promotes the expression of proteins that protect cells from stress. The arginine/lysine transporter, cat-1, is induced during amino acid starvation by transcriptional and post-transcriptional mechanisms. It is shown in the present study that the transient induction of cat-1 transcription is regulated by the stress response pathway that involves phosphorylation of the translation initiation factor, eIF2 (eukaryotic initiation factor-2). This phosphorylation induces expression of the bZIP (basic leucine zipper protein) transcription factors C/EBP (CCAAT/enhancer-binding protein)-beta and ATF (activating transcription factor) 4, which in turn induces ATF3. Transfection experiments in control and mutant cells, and chromatin immunoprecipitations showed that ATF4 activates, whereas ATF3 represses cat-1 transcription, via an AARE (amino acid response element), TGATGAAAC, in the first exon of the cat-1 gene, which functions both in the endogenous and in a heterologous promoter. ATF4 and C/EBPbeta activated transcription when expressed in transfected cells and they bound as heterodimers to the AARE in vitro. The induction of transcription by ATF4 was inhibited by ATF3, which also bound to the AARE as a heterodimer with C/EBPbeta. These results suggest that the transient increase in cat-1 transcription is due to transcriptional activation caused by ATF4 followed by transcriptional repression by ATF3 via a feedback mechanism.
Figures


Similar articles
-
Activation of the ATF3 gene through a co-ordinated amino acid-sensing response programme that controls transcriptional regulation of responsive genes following amino acid limitation.Biochem J. 2007 Jan 1;401(1):299-307. doi: 10.1042/BJ20061261. Biochem J. 2007. PMID: 16989641 Free PMC article.
-
Amino acid deprivation induces the transcription rate of the human asparagine synthetase gene through a timed program of expression and promoter binding of nutrient-responsive basic region/leucine zipper transcription factors as well as localized histone acetylation.J Biol Chem. 2004 Dec 3;279(49):50829-39. doi: 10.1074/jbc.M409173200. Epub 2004 Sep 22. J Biol Chem. 2004. PMID: 15385533
-
Transcriptional control of the arginine/lysine transporter, cat-1, by physiological stress.J Biol Chem. 2003 Dec 12;278(50):50000-9. doi: 10.1074/jbc.M305903200. Epub 2003 Sep 30. J Biol Chem. 2003. PMID: 14523001
-
Coping with stress: eIF2 kinases and translational control.Biochem Soc Trans. 2006 Feb;34(Pt 1):7-11. doi: 10.1042/BST20060007. Biochem Soc Trans. 2006. PMID: 16246168 Review.
-
Activating transcription factor 4.Int J Biochem Cell Biol. 2008;40(1):14-21. doi: 10.1016/j.biocel.2007.01.020. Epub 2007 Jan 28. Int J Biochem Cell Biol. 2008. PMID: 17466566 Review.
Cited by
-
Temporal regulation of Cat-1 (cationic amino acid transporter-1) gene transcription during endoplasmic reticulum stress.Biochem J. 2010 Jul 1;429(1):215-24. doi: 10.1042/BJ20100286. Biochem J. 2010. PMID: 20408811 Free PMC article.
-
Arginine homeostasis and transport in the human pathogen Leishmania donovani.J Biol Chem. 2009 Jul 24;284(30):19800-7. doi: 10.1074/jbc.M901066200. Epub 2009 May 13. J Biol Chem. 2009. PMID: 19439418 Free PMC article.
-
Targeting the Proline-Glutamine-Asparagine-Arginine Metabolic Axis in Amino Acid Starvation Cancer Therapy.Pharmaceuticals (Basel). 2021 Jan 18;14(1):72. doi: 10.3390/ph14010072. Pharmaceuticals (Basel). 2021. PMID: 33477430 Free PMC article. Review.
-
The market for amino acids: understanding supply and demand of substrate for more efficient milk protein synthesis.J Anim Sci Biotechnol. 2020 Nov 12;11(1):108. doi: 10.1186/s40104-020-00514-6. J Anim Sci Biotechnol. 2020. PMID: 33292704 Free PMC article. Review.
-
Differential control of the CCAAT/enhancer-binding protein beta (C/EBPbeta) products liver-enriched transcriptional activating protein (LAP) and liver-enriched transcriptional inhibitory protein (LIP) and the regulation of gene expression during the response to endoplasmic reticulum stress.J Biol Chem. 2008 Aug 15;283(33):22443-56. doi: 10.1074/jbc.M801046200. Epub 2008 Jun 11. J Biol Chem. 2008. PMID: 18550528 Free PMC article.
References
-
- Palacin M., Estevez R., Bertran J., Zorzano A. Molecular biology of mammalian plasma membrane amino acid transporters. Physiol. Rev. 1998;78:969–1054. - PubMed
-
- Hatzoglou M., Fernandez J., Yaman I., Closs E. Regulation of cationic amino acid transport: the story of the CAT-1 transporter. Annu. Rev. Nutr. 2004;24:377–399. - PubMed
-
- Fernandez J., Lopez A. B., Wang C., Mishra R., Zhou L., Yaman I., Snider M. D., Hatzoglou M., Hatzolgou M. Transcriptional control of the arginine/lysine transporter, cat-1, by physiological stress. J. Biol. Chem. 2003;278:50000–50009. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

