TILLING is an effective reverse genetics technique for Caenorhabditis elegans
- PMID: 17049087
- PMCID: PMC1626091
- DOI: 10.1186/1471-2164-7-262
TILLING is an effective reverse genetics technique for Caenorhabditis elegans
Abstract
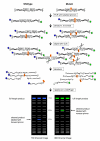
Background: TILLING (Targeting Induced Local Lesions in Genomes) is a reverse genetic technique based on the use of a mismatch-specific enzyme that identifies mutations in a target gene through heteroduplex analysis. We tested this technique in Caenorhabditis elegans, a model organism in which genomics tools have been well developed, but limitations in reverse genetics have restricted the number of heritable mutations that have been identified.
Results: To determine whether TILLING represents an effective reverse genetic strategy for C. elegans we generated an EMS-mutagenised population of approximately 1500 individuals and screened for mutations in 10 genes. A total of 71 mutations were identified by TILLING, providing multiple mutant alleles for every gene tested. Some of the mutations identified are predicted to be silent, either because they are in non-coding DNA or because they affect the third bp of a codon which does not change the amino acid encoded by that codon. However, 59% of the mutations identified are missense alleles resulting in a change in one of the amino acids in the protein product of the gene, and 3% are putative null alleles which are predicted to eliminate gene function. We compared the types of mutation identified by TILLING with those previously reported from forward EMS screens and found that 96% of TILLING mutations were G/C-to-A/T transitions, a rate significantly higher than that found in forward genetic screens where transversions and deletions were also observed. The mutation rate we achieved was 1/293 kb, which is comparable to the mutation rate observed for TILLING in other organisms.
Conclusion: We conclude that TILLING is an effective and cost-efficient reverse genetics tool in C. elegans. It complements other reverse genetic techniques in this organism, can provide an allelic series of mutations for any locus and does not appear to have any bias in terms of gene size or location. For eight of the 10 target genes screened, TILLING has provided the first genetically heritable mutations which can be used to study their functions in vivo.
Figures
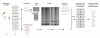


References
-
- Hodgkin J. WormBook. The C. elegans Research Community; 2005. Introduction to genetics and genomics (September 6, 2005)http://www.wormbook.org
-
- Barr MM. Super models. Physiol Genomics. 2003;13:15–24. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

