TLK1B promotes repair of UV-damaged DNA through chromatin remodeling by Asf1
- PMID: 17054786
- PMCID: PMC1626478
- DOI: 10.1186/1471-2199-7-37
TLK1B promotes repair of UV-damaged DNA through chromatin remodeling by Asf1
Abstract
Background: The mammalian protein kinase TLK1 is a homologue of Tousled, a gene involved in flower development in Arabidopsis thaliana. The function of TLK1 is not well known, although knockout of the gene in Drosophila, or expression of a dominant negative mutant in mouse mammary cells causes loss of nuclear divisions and chromosome mis-segregation. TLK1B is a splice variant of TLK1 and it confers radioresistance in a normal mammary mouse cell line possibly due to increased chromatin remodeling capacity, but the mechanism of resistance remains to be fully elucidated.
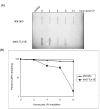
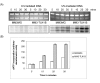
Results: We now show that TLK1B also affords protection against UV radiation. We find that nuclear extracts isolated from TLK1B-containing mouse cells promote more efficient chromatin assembly than comparable extracts lacking TLK1B. TLK1B-containing extracts are also more efficient in repair of UV-damaged plasmid DNA assembled into nucleosomes. One of the two known substrates of TLK1 (or TLK1B) is the histone chaperone Asf1, and immuno-inactivation experiments suggest that TLK1B increases UV-repair through the action of Asf1 on chromatin assembly/disassembly.
Conclusion: Our studies provide evidence for TLK1B-mediated phosphorylation of Asf1 triggering DNA repair. We suggest that this occurs via Asf1-mediated chromatin assembly at the sites of UV damage.
Figures





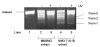

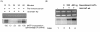

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

