Dissecting the ubiquitin pathway by mass spectrometry
- PMID: 17055348
- PMCID: PMC1828906
- DOI: 10.1016/j.bbapap.2006.09.004
Dissecting the ubiquitin pathway by mass spectrometry
Abstract
Protein modification by ubiquitin is a central regulatory mechanism in eukaryotic cells. Recent proteomics developments in mass spectrometry enable systematic analysis of cellular components in the ubiquitin pathway. Here, we review the advances in analyzing ubiquitinated substrates, determining modified lysine residues, quantifying polyubiquitin chain topologies, as well as profiling deubiquitinating enzymes based on the activity. Moreover, proteomic approaches have been developed for probing the interactome of proteasome and for identifying proteins with ubiquitin-binding domains. Similar strategies have been applied on the studies of the modification by ubiquitin-like proteins as well. These strategies are discussed with respect to their advantages, limitations and potential improvements. While the utilization of current methodologies has rapidly expanded the scope of protein modification by the ubiquitin family, a more active role is anticipated in the functional studies with the emergence of quantitative mass spectrometry.
Figures

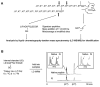
References
-
- Hershko A, Ciechanover A. The ubiquitin system. Annu Rev Biochem. 1998;67:425–79. - PubMed
-
- Varshavsky A. Regulated protein degradation. Trends Biochem Sci. 2005;30:283–6. - PubMed
-
- Weissman AM. Themes and variations on ubiquitylation. Nat Rev Mol Cell Biol. 2001;2:169–78. - PubMed
-
- Hicke L, Dunn R. Regulation of membrane protein transport by ubiquitin and ubiquitin-binding proteins. Annu Rev Cell Dev Biol. 2003;19:141–72. - PubMed
-
- Pickart CM. Mechanisms underlying ubiquitination. Annu Rev Biochem. 2001;70:503–33. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

