Cloning and expression of the Erwinia carotovora subsp. carotovora gene encoding the low-molecular-weight bacteriocin carocin S1
- PMID: 17071754
- PMCID: PMC1797388
- DOI: 10.1128/JB.01090-06
Cloning and expression of the Erwinia carotovora subsp. carotovora gene encoding the low-molecular-weight bacteriocin carocin S1
Abstract
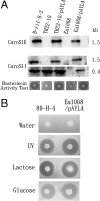
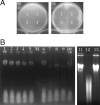
The purpose of this study was to clone the carocin S1 gene and express it in a non-carocin-producing strain of Erwinia carotovora. A mutant, TH22-10, which produced a high-molecular-weight bacteriocin but not a low-molecular-weight bacteriocin, was obtained by Tn5 insertional mutagenesis using H-rif-8-2 (a spontaneous rifampin-resistant mutant of Erwinia carotovora subsp. carotovora 89-H-4). Using thermal asymmetric interlaced PCR, the DNA sequence from the Tn5 insertion site and the DNA sequence of the contiguous 2,280-bp region were determined. Two complete open reading frames (ORF), designated ORF2 and ORF3, were identified within the sequence fragment. ORF2 and ORF3 were identified with the carocin S1 genes, caroS1K (ORF2) and caroS1I (ORF3), which, respectively, encode a killing protein (CaroS1K) and an immunity protein (CaroS1I). These genes were homologous to the pyocin S3 gene and the pyocin AP41 gene. Carocin S1 was expressed in E. carotovora subsp. carotovora Ea1068 and replicated in TH22-10 but could not be expressed in Escherichia coli (JM101) because a consensus sequence resembling an SOS box was absent. A putative sequence similar to the consensus sequence for the E. coli cyclic AMP receptor protein binding site (-312 bp) was found upstream of the start codon. Production of this bacteriocin was also induced by glucose and lactose. The homology search results indicated that the carocin S1 gene (between bp 1078 and bp 1704) was homologous to the pyocin S3 and pyocin AP41 genes in Pseudomonas aeruginosa. These genes encode proteins with nuclease activity (domain 4). This study found that carocin S1 also has nuclease activity.
Figures


Similar articles
-
Identification and cloning of an Erwinia carotovora subsp. carotovora bacteriocin regulator gene by insertional mutagenesis.J Bacteriol. 1999 Mar;181(6):1953-7. doi: 10.1128/JB.181.6.1953-1957.1999. J Bacteriol. 1999. PMID: 10074096 Free PMC article.
-
Identification and characterization of the bacteriocin Carocin S3 from the multiple bacteriocin producing strain of Pectobacterium carotovorum subsp. carotovorum.BMC Microbiol. 2020 Sep 1;20(1):273. doi: 10.1186/s12866-020-01955-9. BMC Microbiol. 2020. PMID: 32867691 Free PMC article.
-
Cloning, purification, and functional characterization of Carocin S2, a ribonuclease bacteriocin produced by Pectobacterium carotovorum.BMC Microbiol. 2011 May 12;11:99. doi: 10.1186/1471-2180-11-99. BMC Microbiol. 2011. PMID: 21569432 Free PMC article.
-
The pyocins of Pseudomonas aeruginosa.Biochimie. 2002 May-Jun;84(5-6):499-510. doi: 10.1016/s0300-9084(02)01422-0. Biochimie. 2002. PMID: 12423794 Review.
-
Turning Over a New Leaf: Bacteriocins Going Green.Trends Microbiol. 2018 Jan;26(1):1-2. doi: 10.1016/j.tim.2017.11.001. Epub 2017 Nov 14. Trends Microbiol. 2018. PMID: 29150081 Review.
Cited by
-
A Novel Deoxyribonuclease Low-Molecular-Weight Bacteriocin, Carocin S4, from Pectobacterium carotovorum subsp. carotovorum.Microorganisms. 2023 Jul 22;11(7):1854. doi: 10.3390/microorganisms11071854. Microorganisms. 2023. PMID: 37513026 Free PMC article.
-
New type of antimicrobial protein produced by the plant pathogen Clavibacter michiganensis subsp. michiganensis.Appl Environ Microbiol. 2013 Sep;79(18):5721-7. doi: 10.1128/AEM.01065-13. Epub 2013 Jul 12. Appl Environ Microbiol. 2013. PMID: 23851100 Free PMC article.
-
The crystal structure of the lipid II-degrading bacteriocin syringacin M suggests unexpected evolutionary relationships between colicin M-like bacteriocins.J Biol Chem. 2012 Nov 9;287(46):38876-88. doi: 10.1074/jbc.M112.400150. Epub 2012 Sep 20. J Biol Chem. 2012. PMID: 22995910 Free PMC article.
-
Comparative genomics of plant-associated Pseudomonas spp.: insights into diversity and inheritance of traits involved in multitrophic interactions.PLoS Genet. 2012 Jul;8(7):e1002784. doi: 10.1371/journal.pgen.1002784. Epub 2012 Jul 5. PLoS Genet. 2012. PMID: 22792073 Free PMC article.
-
Intercepting biological messages: Antibacterial molecules targeting nucleic acids during interbacterial conflicts.Genet Mol Biol. 2023 Mar 6;46(1 Suppl 2):e20220266. doi: 10.1590/1678-4685-GMB-2022-0266. eCollection 2023. Genet Mol Biol. 2023. PMID: 36880694 Free PMC article.
References
-
- Bolivar, F., R. L. Rodriguez, P. J. Greene, M. C. Betlach, H. L. Heyneker, and H. W. Boyer. 1977. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene 2:95-113. - PubMed
-
- Chuang, D.-Y., Y. Gunji, A. G. Kyeremeh, Y. Takahara, and T. Kikumoto. 1998. Cloning of bacteriocin regulator gene (brg) from Erwinia carotovora subsp. carotovora, abstr. 251, p. 389-390. Abstr. Annu. Meet. Soc. 1998. Phytopathological Society of Japan, Sapporo, Japan. (In Japanese.)
-
- Duport, C., C. Baysse, and Y. Michel-Briand. 1995. Molecular characterization of pyocin S3, a novel S-type pyocin from Pseudomonas aeruginosa. J. Biol. Chem. 270:8920-8927. - PubMed
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

