Fluorescence correlation spectroscopy shows that monomeric polyglutamine molecules form collapsed structures in aqueous solutions
- PMID: 17075061
- PMCID: PMC1629004
- DOI: 10.1073/pnas.0608175103
Fluorescence correlation spectroscopy shows that monomeric polyglutamine molecules form collapsed structures in aqueous solutions
Abstract
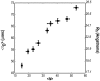
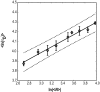
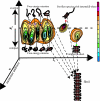
We have used fluorescence correlation spectroscopy measurements to quantify the hydrodynamic sizes of monomeric polyglutamine as a function of chain length (N) by measuring the scaling of translational diffusion times (tau(D)) for the peptide series (Gly)-(Gln)(N)-Cys-Lys(2) in aqueous solution. We find that tau(D) scales with N as tau(o)N(nu) and therefore ln(tau(D)) = ln(tau(o)) + nuln(N). The values for nu and ln(tau(o)) are 0.32 +/- 0.02 and 3.04 +/- 0.08, respectively. Based on these observations, we conclude that water is a polymeric poor solvent for polyglutamine. Previous studies have shown that monomeric polyglutamine is intrinsically disordered. These observations combined with our fluorescence correlation spectroscopy data suggest that the ensemble for monomeric polyglutamine is made up of a heterogeneous collection of collapsed structures. This result is striking because the preference for collapsed structures arises despite the absence of residues deemed to be hydrophobic in the sequence constructs studied. Working under the assumption that the driving forces for collapse are similar to those for aggregation, we discuss the implications of our results for the thermodynamics and kinetics of polyglutamine aggregation, a process that has been implicated in the molecular mechanism of Huntington's disease.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Atomistic simulations of the effects of polyglutamine chain length and solvent quality on conformational equilibria and spontaneous homodimerization.J Mol Biol. 2008 Dec 5;384(1):279-97. doi: 10.1016/j.jmb.2008.09.026. Epub 2008 Sep 18. J Mol Biol. 2008. PMID: 18824003 Free PMC article.
-
Quantitative characterization of intrinsic disorder in polyglutamine: insights from analysis based on polymer theories.Biophys J. 2007 Sep 15;93(6):1923-37. doi: 10.1529/biophysj.107.110080. Epub 2007 May 25. Biophys J. 2007. PMID: 17526581 Free PMC article.
-
Modulation of polyglutamine conformations and dimer formation by the N-terminus of huntingtin.J Mol Biol. 2010 Mar 12;396(5):1295-309. doi: 10.1016/j.jmb.2009.12.017. Epub 2009 Dec 21. J Mol Biol. 2010. PMID: 20026071 Free PMC article.
-
Examining polyglutamine peptide length: a connection between collapsed conformations and increased aggregation.J Mol Biol. 2009 Nov 6;393(4):978-92. doi: 10.1016/j.jmb.2009.08.034. Epub 2009 Aug 20. J Mol Biol. 2009. PMID: 19699209 Free PMC article.
-
Polyglutamine aggregation behavior in vitro supports a recruitment mechanism of cytotoxicity.J Mol Biol. 2001 Aug 3;311(1):173-82. doi: 10.1006/jmbi.2001.4850. J Mol Biol. 2001. PMID: 11469866
Cited by
-
Understanding protein aggregation from the view of monomer dynamics.Mol Biosyst. 2013 Jan 27;9(1):29-35. doi: 10.1039/c2mb25334h. Epub 2012 Oct 26. Mol Biosyst. 2013. PMID: 23104145 Free PMC article. Review.
-
Describing sequence-ensemble relationships for intrinsically disordered proteins.Biochem J. 2013 Jan 15;449(2):307-18. doi: 10.1042/BJ20121346. Biochem J. 2013. PMID: 23240611 Free PMC article. Review.
-
Dynamic imaging by fluorescence correlation spectroscopy identifies diverse populations of polyglutamine oligomers formed in vivo.J Biol Chem. 2012 Jul 27;287(31):26136-45. doi: 10.1074/jbc.M112.362764. Epub 2012 Jun 5. J Biol Chem. 2012. PMID: 22669943 Free PMC article.
-
Conformation sensors that distinguish monomeric proteins from oligomers in live cells.Chem Biol. 2010 Apr 23;17(4):371-9. doi: 10.1016/j.chembiol.2010.03.011. Chem Biol. 2010. PMID: 20416508 Free PMC article.
-
SWI/SNF senses carbon starvation with a pH-sensitive low-complexity sequence.Elife. 2022 Feb 7;11:e70344. doi: 10.7554/eLife.70344. Elife. 2022. PMID: 35129437 Free PMC article.
References
-
- Monaco S, Zanusso G, Mazzucco S, Rizzuto N. Curr Med Chem. 2006;13:1903–1913. - PubMed
-
- Obici L, Perfetti V, Palladini G, Moratti R, Merlini G. Biochim Biophys Acta. 2005;1753:11–22. - PubMed
-
- Westermark P. FEBS J. 2005;272:5942–5949. - PubMed
-
- Dobson CM. Semin Cell Dev Biol. 2004;15:3–16. - PubMed
-
- Uversky VN, Fink AL. Biochim Biophys Acta. 2004;1698:131–153. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

