Role of metal ions in catalysis by HIV integrase analyzed using a quantitative PCR disintegration assay
- PMID: 17085478
- PMCID: PMC1693899
- DOI: 10.1093/nar/gkl862
Role of metal ions in catalysis by HIV integrase analyzed using a quantitative PCR disintegration assay
Abstract
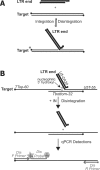
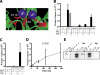
Paired metal ions have been proposed to be central to the catalytic mechanisms of RNase H nucleases, bacterial transposases, Holliday junction resolvases, retroviral integrases and many other enzymes. Here we present a sensitive assay for DNA transesterification in which catalysis by human immunodeficiency virus-type 1 (HIV-1) integrase (IN) connects two DNA strands (disintegration reaction), allowing detection using quantitative PCR (qPCR). We present evidence suggesting that the three acidic residues of the IN active site function through metal binding using metal rescue. In this method, the catalytic acidic residues were each substituted with cysteines. Mn2+ binds tightly to the sulfur atoms of the cysteine residues, but Mg2+ does not. We found that Mn2+, but not Mg2+, could rescue catalysis of each cysteine-substituted enzyme, providing evidence for functionally important metal binding by all three residues. We also used the PCR-boosted assay to show that HIV-1 IN could carry out transesterification reactions involving DNA 5' hydroxyl groups as well as 3' hydroxyls as nucleophiles. Lastly, we show that Mn2+ by itself (i.e. without enzyme) can catalyze formation of a low level of PCR-amplifiable product under extreme conditions, allowing us to estimate the rate enhancement due to the IN-protein scaffold as at least 60 million-fold.
Figures


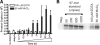
References
-
- Pan T., Long D.M., Uhlenbeck O.C. Divalent metal ions in RNA folding and catalysis. In: Gesteland R.F., Atkins J.F., editors. The RNA World. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1993. pp. 271–302.
-
- Rice P., Craigie R., Davies D.R. Retroviral integrases and their cousins. Curr. Opin. Struct. Biol. 1996;6:76–83. - PubMed
-
- Yang W., Steitz T.A. Recombining the structures of HIV integrase, RuvC, and RNase H. Structure. 1995;3:131–134. - PubMed

