The cPLA2 C2alpha domain in solution: structure and dynamics of its Ca2+-activated and cation-free states
- PMID: 17085504
- PMCID: PMC1779961
- DOI: 10.1529/biophysj.106.091850
The cPLA2 C2alpha domain in solution: structure and dynamics of its Ca2+-activated and cation-free states
Abstract
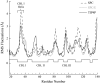
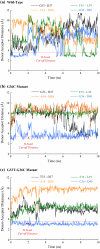
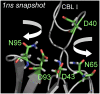
Cytosolic phospholipase A2 is involved in several signal transduction pathways where it catalyses release of arachidonic acid from intracellular lipid membranes. Its membrane insertion is facilitated by its independently folding C2alpha domain, which is activated by the binding of two intracellular Ca2+ ions. However, the details of its membrane-insertion mechanism, including its Ca2+-activation mechanism, are not understood. There are several unresolved issues, including the following. There are two experimentally resolved structures of the Ca2+-activated state of its isolated C2alpha domain, one determined using x-ray crystallography and the other determined using NMR spectroscopy, which differ from each other significantly in the spatial region that inserts into the membrane. This by itself adds to ambiguities associated with investigations targeting its mechanism of membrane insertion. Furthermore, there is no experimentally determined structure of its cation-free state, which hinders investigations associated with its cation-activation mechanism. In this work, we generate several unrestrained molecular dynamics trajectories of its isolated C2alpha domain in solution (equivalent to approximately 60 ns) and investigate these issues. Our main results are as follows: a), the Ca2+ coordination scheme of the domain is consistent with the x-ray structure and with previous mutagenesis studies; b), the helical segment of the Ca2+-binding loop, CBL-I, undergoes nanosecond timescale flexing (but not an unwinding), as can be inferred from physiological temperature NMR data and in contrast to low temperature x-ray data; and c), removal of the two activating Ca2+ ions from their binding pockets does not alter the backbone structure of the domain, a result consistent with electron paramagnetic resonance data.
Figures






Similar articles
-
Self-induced docking site of a deeply embedded peripheral membrane protein.Biophys J. 2007 Jan 15;92(2):517-24. doi: 10.1529/biophysj.106.090704. Epub 2006 Oct 27. Biophys J. 2007. PMID: 17071664 Free PMC article.
-
A structure-function study of the C2 domain of cytosolic phospholipase A2. Identification of essential calcium ligands and hydrophobic membrane binding residues.J Biol Chem. 1999 Apr 2;274(14):9665-72. doi: 10.1074/jbc.274.14.9665. J Biol Chem. 1999. PMID: 10092653
-
Ca2+-signaling cycle of a membrane-docking C2 domain.Biochemistry. 1997 Oct 7;36(40):12011-8. doi: 10.1021/bi9717340. Biochemistry. 1997. PMID: 9340010
-
Cytosolic phospholipase A2.J Lipid Mediat Cell Signal. 1995 Oct;12(2-3):83-117. doi: 10.1016/0929-7855(95)00012-f. J Lipid Mediat Cell Signal. 1995. PMID: 8777586 Review.
-
Structure, function and regulation of Ca2+-sensitive cytosolic phospholipase A2 (cPLA2).FEBS Lett. 1997 Jun 23;410(1):49-53. doi: 10.1016/s0014-5793(97)00322-0. FEBS Lett. 1997. PMID: 9247121 Review.
Cited by
-
Structural transitions in ion coordination driven by changes in competition for ligand binding.J Am Chem Soc. 2008 Nov 19;130(46):15405-19. doi: 10.1021/ja803575y. Epub 2008 Oct 28. J Am Chem Soc. 2008. PMID: 18954053 Free PMC article.
-
K+/Na+ selectivity in K channels and valinomycin: over-coordination versus cavity-size constraints.J Mol Biol. 2008 Feb 8;376(1):13-22. doi: 10.1016/j.jmb.2007.11.059. Epub 2007 Nov 28. J Mol Biol. 2008. PMID: 18155244 Free PMC article.
-
Structural and activity characterization of human PHPT1 after oxidative modification.Sci Rep. 2016 Apr 1;6:23658. doi: 10.1038/srep23658. Sci Rep. 2016. PMID: 27034094 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

