Genomic structure and phylogeny of the plant pathogen Ralstonia solanacearum inferred from gene distribution analysis
- PMID: 17085551
- PMCID: PMC1797399
- DOI: 10.1128/JB.00999-06
Genomic structure and phylogeny of the plant pathogen Ralstonia solanacearum inferred from gene distribution analysis
Abstract
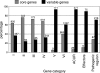
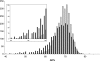
In the present study, we investigated the gene distribution among strains of the highly polymorphic plant pathogenic beta-proteobacterium Ralstonia solanacearum, paying particular attention to the status of known or candidate pathogenicity genes. Based on the use of comparative genomic hybridization on a pangenomic microarray for the GMI1000 reference strain, we have defined the conditions that allowed comparison of the repertoires of genes among a collection of 18 strains that are representative of the biodiversity of the R. solanacearum species. This identified a list of 2,690 core genes present in all tested strains. As a corollary, a list of 2,338 variable genes within the R. solanacearum species has been defined. The hierarchical clustering based on the distribution of variable genes is fully consistent with the phylotype classification that was previously defined from the nucleotide sequence analysis of four genes. The presence of numerous pathogenicity-related genes in the core genome indicates that R. solanacearum is an ancestral pathogen. The results establish the long coevolution of the two replicons that constitute the bacterial genome. We also demonstrate the clustering of variable genes in genomic islands. Most genomic islands are included in regions with an alternative codon usage, suggesting that they originate from acquisition of foreign genes through lateral gene transfers. Other genomic islands correspond to genes that have the same base composition as core genes, suggesting that they either might be ancestral genes lost by deletion in certain strains or might originate from horizontal gene transfers.
Figures



Similar articles
-
Genomes of three tomato pathogens within the Ralstonia solanacearum species complex reveal significant evolutionary divergence.BMC Genomics. 2010 Jun 15;11:379. doi: 10.1186/1471-2164-11-379. BMC Genomics. 2010. PMID: 20550686 Free PMC article.
-
Distribution and sequence analysis of a family of type ill-dependent effectors correlate with the phylogeny of Ralstonia solanacearum strains.Mol Plant Microbe Interact. 2004 Aug;17(8):931-40. doi: 10.1094/MPMI.2004.17.8.931. Mol Plant Microbe Interact. 2004. PMID: 15305614
-
Ralstonia solanacearum virulence increased following large interstrain gene transfers by natural transformation.Mol Plant Microbe Interact. 2011 Apr;24(4):497-505. doi: 10.1094/MPMI-09-10-0197. Mol Plant Microbe Interact. 2011. PMID: 21190441
-
Pathogenomics of the Ralstonia solanacearum species complex.Annu Rev Phytopathol. 2012;50:67-89. doi: 10.1146/annurev-phyto-081211-173000. Epub 2012 May 1. Annu Rev Phytopathol. 2012. PMID: 22559068 Review.
-
Ralstonia solanacearum, a widespread bacterial plant pathogen in the post-genomic era.Mol Plant Pathol. 2013 Sep;14(7):651-62. doi: 10.1111/mpp.12038. Epub 2013 May 30. Mol Plant Pathol. 2013. PMID: 23718203 Free PMC article. Review.
Cited by
-
A unique DNA repair and recombination gene (recN) sequence for identification and intraspecific molecular typing of bacterial wilt pathogen Ralstonia solanacearum and its comparative analysis with ribosomal DNA sequences.J Biosci. 2013 Jun;38(2):267-78. doi: 10.1007/s12038-013-9312-0. J Biosci. 2013. PMID: 23660661
-
Biosynthesis of a complex yersiniabactin-like natural product via the mic locus in phytopathogen Ralstonia solanacearum.Appl Environ Microbiol. 2011 Sep;77(17):6117-24. doi: 10.1128/AEM.05198-11. Epub 2011 Jul 1. Appl Environ Microbiol. 2011. PMID: 21724891 Free PMC article.
-
Type three effector gene distribution and sequence analysis provide new insights into the pathogenicity of plant-pathogenic Xanthomonas arboricola.Appl Environ Microbiol. 2012 Jan;78(2):371-84. doi: 10.1128/AEM.06119-11. Epub 2011 Nov 18. Appl Environ Microbiol. 2012. PMID: 22101042 Free PMC article.
-
The ecological coherence of high bacterial taxonomic ranks.Nat Rev Microbiol. 2010 Jul;8(7):523-9. doi: 10.1038/nrmicro2367. Nat Rev Microbiol. 2010. PMID: 20531276 Review.
-
Comparative genomics reveals diversity among xanthomonads infecting tomato and pepper.BMC Genomics. 2011 Mar 11;12:146. doi: 10.1186/1471-2164-12-146. BMC Genomics. 2011. PMID: 21396108 Free PMC article.
References
-
- Alfano, J. R., and A. Collmer. 2004. Type III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu. Rev. Phytopathol. 42:385-414. - PubMed
-
- Bertolla, F., A. Frostegård, B. Brito, X. Nesme, and P. Simonet. 1999. During infection of its hosts, the plant pathogen Ralstonia solanacearum naturally develops a state of competence and exchanges genetic material. Mol. Plant-Microbe Interact. 12:467-472.
-
- Boucher, C., A. Martinel, P. Barberis, G. Alloing, and C. Zischek. 1985. Virulence genes are carried by a megaplasmid of the plant pathogen Pseudomonas solanacearum. Mol. Gen. Genet. 205:270-275.
-
- Boureau, T., H. El Maarouf-Bouteau, A. Garnier, M. N. Brisset, C. Perino, I. Pucheu, and M. A. Barny. 2006. DspA/E, a type III effector essential for Erwinia amylovora pathogenicity and growth in planta, induces cell death in host apple and nonhost tobacco plants. Mol. Plant-Microbe Interact. 19:16-24. - PubMed
-
- Burrus, V., and M. K. Waldor. 2004. Shaping bacterial genomes with integrative and conjugative elements. Res. Microbiol. 155:376-386. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

