Extent of horizontal gene transfer in evolution of Streptococci of the salivarius group
- PMID: 17085557
- PMCID: PMC1797340
- DOI: 10.1128/JB.01058-06
Extent of horizontal gene transfer in evolution of Streptococci of the salivarius group
Abstract
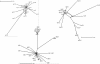
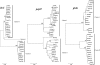
The phylogenetically closely related species Streptococcus salivarius and Streptococcus vestibularis are oral bacteria that are considered commensals, although they can also be found in human infections. The relationship between these two species and the relationship between strains isolated from carriers and strains responsible for invasive infections were investigated by multilocus sequence typing and additional sequence analysis. The clustering of several S. vestibularis alleles and the extent of genomic divergence at certain loci support the conclusion that S. salivarius and S. vestibularis are separate species. The level of sequence diversity in S. salivarius alleles is generally high, whereas that in S. vestibularis alleles is low at certain loci, indicating that the latter species might have evolved recently. Cluster analysis indicated that there has been genetic exchange between S. salivarius and S. vestibularis at three of the nine loci investigated. Horizontal gene transfer between streptococci belonging to the S. salivarius group and other oral streptococci was also detected at several loci. A high level of recombination in S. salivarius was revealed by allele index association and split decomposition sequence analyses. Commensal and infection-associated S. salivarius strains could not be distinguished by cluster analysis, suggesting that the pathogen isolates are opportunistic. Taken together, our results indicate that there is a high level of gene exchange that contributes to the evolution of two streptococcal species from the human oral cavity.
Figures




Similar articles
-
Genomics of Streptococcus salivarius, a major human commensal.Infect Genet Evol. 2015 Jul;33:381-92. doi: 10.1016/j.meegid.2014.10.001. Epub 2014 Oct 13. Infect Genet Evol. 2015. PMID: 25311532 Review.
-
Evolutionary relationships among salivarius streptococci as inferred from multilocus phylogenies based on 16S rRNA-encoding, recA, secA, and secY gene sequences.BMC Microbiol. 2009 Oct 30;9:232. doi: 10.1186/1471-2180-9-232. BMC Microbiol. 2009. PMID: 19878555 Free PMC article.
-
Diversity of Integrative and Conjugative Elements of Streptococcus salivarius and Their Intra- and Interspecies Transfer.Appl Environ Microbiol. 2017 Jun 16;83(13):e00337-17. doi: 10.1128/AEM.00337-17. Print 2017 Jul 1. Appl Environ Microbiol. 2017. PMID: 28432093 Free PMC article.
-
A New Species-Specific Typing Method for Salivarius Group Streptococci Based on the Dephospho-Coenzyme A Kinase (coaE) Gene Sequencing.Front Cell Infect Microbiol. 2021 Aug 6;11:685657. doi: 10.3389/fcimb.2021.685657. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 34422679 Free PMC article.
-
Genomics, evolution, and molecular epidemiology of the Streptococcus bovis/Streptococcus equinus complex (SBSEC).Infect Genet Evol. 2015 Jul;33:419-36. doi: 10.1016/j.meegid.2014.09.017. Epub 2014 Sep 16. Infect Genet Evol. 2015. PMID: 25233845 Review.
Cited by
-
Evidence for niche adaptation in the genome of the bovine pathogen Streptococcus uberis.BMC Genomics. 2009 Jan 28;10:54. doi: 10.1186/1471-2164-10-54. BMC Genomics. 2009. PMID: 19175920 Free PMC article.
-
Multilocus sequence typing of Streptococcus thermophilus from naturally fermented dairy foods in China and Mongolia.BMC Microbiol. 2015 Oct 26;15:236. doi: 10.1186/s12866-015-0551-0. BMC Microbiol. 2015. PMID: 26497818 Free PMC article.
-
Comparative genome analysis of Streptococcus infantarius subsp. infantarius CJ18, an African fermented camel milk isolate with adaptations to dairy environment.BMC Genomics. 2013 Mar 22;14:200. doi: 10.1186/1471-2164-14-200. BMC Genomics. 2013. PMID: 23521820 Free PMC article.
-
Insight into the evolution of the histidine triad protein (HTP) family in Streptococcus.PLoS One. 2013;8(3):e60116. doi: 10.1371/journal.pone.0060116. Epub 2013 Mar 20. PLoS One. 2013. PMID: 23527301 Free PMC article.
-
Complete genome sequence of the pigmented Streptococcus thermophilus strain JIM8232.J Bacteriol. 2011 Oct;193(19):5581-2. doi: 10.1128/JB.05404-11. J Bacteriol. 2011. PMID: 21914889 Free PMC article.
References
-
- Bolotin, A., B. Quinquis, P. Renault, A. Sorokin, S. D. Ehrlich, S. Kulakauskas, A. Lapidus, E. Goltsman, M. Mazur, G. D. Pusch, M. Fonstein, R. Overbeek, N. Kyprides, B. Purnelle, D. Prozzi, K. Ngui, D. Masuy, F. Hancy, S. Burteau, M. Boutry, J. Delcour, A. Goffeau, and P. Hols. 2004. Complete sequence and comparative genome analysis of the dairy bacterium Streptococcus thermophilus. Nat. Biotechnol. 22:1554-1558. - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

