Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic
- PMID: 17085596
- PMCID: PMC1859917
- DOI: 10.1073/pnas.0606088103
Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic
Abstract
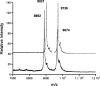
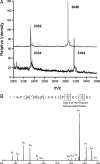
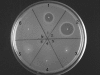
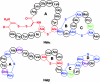
Lantibiotics are ribosomally synthesized peptides that undergo posttranslational modifications to their mature, antimicrobial form. They are characterized by the unique amino acids lanthionine and methyllanthionine, introduced by means of dehydration of Ser/Thr residues followed by reaction of the resulting dehydro amino acids with cysteines to form thioether linkages. Two-component lantibiotics use two peptides that are each posttranslationally modified to yield two functionally distinct products that act in synergy to provide bactericidal activity. By using genetic data instead of isolation, a two-component lantibiotic, haloduracin, was identified in the genome of the Gram-positive alkaliphilic bacterium Bacillus halodurans C-125. We show that heterologously expressed and purified precursor peptides HalA1 and HalA2 are processed by the purified modification enzymes HalM1 and HalM2 in an in vitro reconstitution of the biosynthesis of a two-component lantibiotic. The activity of each HalM enzyme is substrate-specific, and the assay products exhibit antimicrobial activity after removal of their leader sequences at an engineered Factor Xa cleavage site, indicating that correct thioether formation has occurred. Haloduracin's biological activity depends on the presence of both modified peptides. The structures of the two mature haloduracin peptides Halalpha and Halbeta were investigated, indicating that they have similarities as well as some distinct differences compared with other two-component lantibiotics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Delves-Broughton J, Blackburn P, Evans RJ, Hugenholtz J. Antonie van Leeuwenhoek. 1996;69:193–202. - PubMed
-
- Cotter PD, Hill C, Ross RP. Nat Rev Microbiol. 2005;3:777–788. - PubMed
-
- Chatterjee C, Paul M, Xie L, van der Donk WA. Chem Rev. 2005;105:633–684. - PubMed
-
- Schnell N, Entian K-D, Schneider U, Götz F, Zahner H, Kellner R, Jung G. Nature. 1988;333:276–278. - PubMed
-
- van der Meer JR, Rollema HS, Siezen RJ, Beerthuyzen MM, Kuipers OP, de Vos WM. J Biol Chem. 1994;269:3555–3562. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

