Genetic analysis of bacteriocin 43 of vancomycin-resistant Enterococcus faecium
- PMID: 17088377
- PMCID: PMC1636183
- DOI: 10.1128/AEM.00934-06
Genetic analysis of bacteriocin 43 of vancomycin-resistant Enterococcus faecium
Abstract
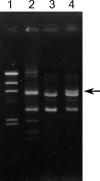
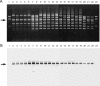
A total of 636 vancomycin-resistant Enterococcus faecium (VRE) isolates obtained between 1994 and 1999 from the Medical School Hospital of the University of Michigan were tested for bacteriocin production. Of the 277 (44%) bacteriocinogenic strains, 21 were active against E. faecalis, E. faecium, E. hirae, E. durans, and Listeria monocytogenes. Of those 21 strains, a representative bacteriocin of strain VRE82, designated bacteriocin 43, was found to be encoded on mobilizable plasmid pDT1 (6.2 kbp). Nine open reading frames (ORFs), ORF1 to ORF9, were presented on pDT1 and were oriented in the same direction. The bacteriocin 43 locus (bac43) consists of the bacteriocin gene bacA (ORF1) and the immunity gene bacB (ORF2). The deduced bacA product is 74 amino acids in length with a putative signal peptide of 30 amino acids at the N terminus. The bacB gene encodes a deduced 95-amino-acid protein without a signal sequence. The predicted mature BacA protein (44 amino acids) showed sequence homology with the membrane-active class IIa bacteriocins of lactic acid bacteria and showed 86% homology with bacteriocin 31 from E. faecalis YI717 and 98% homology with bacteriocin RC714. Southern analysis with a bac43 probe of each plasmid DNA from the 21 strains showed hybridization to a specific fragment corresponding to the 6.2-kbp EcoRI fragment, suggesting that the strains harbored the pDT1-like plasmid (6.2 kb) which encoded the bacteriocin 43-type bacteriocin. The bac43 determinant was not identified among non-VRE clinical isolates.
Figures


References
-
- Balla, E., and L. M. T. Dicks. 2005. Molecular analysis of the gene cluster involved in the production and secretion of enterocins 1071A and 1071B and of the genes responsible for the replication and transfer of plasmid pEF1071. Int. J. Food Microbiol. 99:33-45. - PubMed
-
- Casaus, P., T. Nilsen, L. M. Cintas, I. F. Nes, P. E. Hernandez, and H. Holo. 1997. Enterocin B, a new bacteriocin from Enterococcus faecium T136 which can act synergistically with enterocin A. Microbiology 143:2287-2294. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

