Proteolysis restricts localization of CID, the centromere-specific histone H3 variant of Drosophila, to centromeres
- PMID: 17090596
- PMCID: PMC1693906
- DOI: 10.1093/nar/gkl902
Proteolysis restricts localization of CID, the centromere-specific histone H3 variant of Drosophila, to centromeres
Abstract
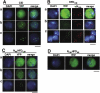
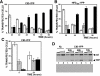
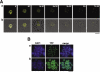
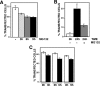
Centromere identity is determined by the formation of a specialized chromatin structure containing the centromere-specific histone H3 variant CENP-A. The precise molecular mechanism(s) accounting for the specific deposition of CENP-A at centromeres are still poorly understood. Centromeric deposition of CENP-A, which is independent of DNA replication, might involve specific chromatin assembly complexes and/or specific interactions with kinetochore components. However, transiently expressed CENP-A incorporates throughout chromatin indicating that CENP-A nucleosomes can also be promiscuously deposited during DNA replication. Therefore, additional mechanisms must exist to prevent deposition of CENP-A nucleosomes during replication and/or to remove them afterwards. Here, using transient expression experiments performed in Drosophila Kc cells, we show that proteasome-mediated degradation restricts localization of Drosophila CENP-A (CID) to centromeres by eliminating mislocalized CID as well as by regulating available CID levels. Regulating available CID levels appears essential to ensure centromeric deposition of transiently expressed CID as, when expression is increased in the presence of proteasome inhibitors, newly synthesized CID mislocalizes. Mislocalization of CID affects cell cycle progression as a high percentage of cells showing mislocalized CID are reactive against alphaPSer(10)H3 antibodies, enter mitosis at a very low frequency and show strong segregation defects. However, cells showing reduced amounts of mislocalized CID show normal cell cycle progression.
Figures


Similar articles
-
Detrimental incorporation of excess Cenp-A/Cid and Cenp-C into Drosophila centromeres is prevented by limiting amounts of the bridging factor Cal1.J Cell Sci. 2010 Nov 1;123(Pt 21):3768-79. doi: 10.1242/jcs.067934. Epub 2010 Oct 12. J Cell Sci. 2010. PMID: 20940262
-
A role of the Trx-G complex in Cid/CENP-A deposition at Drosophila melanogaster centromeres.Chromosoma. 2019 Dec;128(4):503-520. doi: 10.1007/s00412-019-00711-x. Epub 2019 Jun 16. Chromosoma. 2019. PMID: 31203392
-
Centromere identity in Drosophila is not determined in vivo by replication timing.J Cell Biol. 2001 Aug 20;154(4):683-90. doi: 10.1083/jcb.200103001. J Cell Biol. 2001. PMID: 11514585 Free PMC article.
-
Molecular underpinnings of centromere identity and maintenance.Trends Biochem Sci. 2012 Jun;37(6):220-9. doi: 10.1016/j.tibs.2012.01.003. Epub 2012 Mar 10. Trends Biochem Sci. 2012. PMID: 22410197 Free PMC article. Review.
-
Centromere regulation: new players, new rules, new questions.Eur J Cell Biol. 2011 Oct;90(10):805-10. doi: 10.1016/j.ejcb.2011.04.016. Eur J Cell Biol. 2011. PMID: 21684630 Review.
Cited by
-
Haploid Bio-Induction in Plant through Mock Sexual Reproduction.iScience. 2020 Jul 24;23(7):101279. doi: 10.1016/j.isci.2020.101279. Epub 2020 Jun 16. iScience. 2020. PMID: 32619703 Free PMC article.
-
CENP-A nucleosomes localize to transcription factor hotspots and subtelomeric sites in human cancer cells.Epigenetics Chromatin. 2015 Jan 13;8:2. doi: 10.1186/1756-8935-8-2. eCollection 2015. Epigenetics Chromatin. 2015. PMID: 25788983 Free PMC article.
-
The unconventional structure of centromeric nucleosomes.Chromosoma. 2012 Aug;121(4):341-52. doi: 10.1007/s00412-012-0372-y. Epub 2012 May 3. Chromosoma. 2012. PMID: 22552438 Free PMC article. Review.
-
Heterochromatin and RNAi regulate centromeres by protecting CENP-A from ubiquitin-mediated degradation.PLoS Genet. 2018 Aug 8;14(8):e1007572. doi: 10.1371/journal.pgen.1007572. eCollection 2018 Aug. PLoS Genet. 2018. PMID: 30089114 Free PMC article.
-
New clues to understand how CENP-A maintains centromere identity.Cell Div. 2011 May 9;6(1):11. doi: 10.1186/1747-1028-6-11. Cell Div. 2011. PMID: 21554702 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

