Selection for unequal densities of sigma70 promoter-like signals in different regions of large bacterial genomes
- PMID: 17096598
- PMCID: PMC1635534
- DOI: 10.1371/journal.pgen.0020185
Selection for unequal densities of sigma70 promoter-like signals in different regions of large bacterial genomes
Abstract
The evolutionary processes operating in the DNA regions that participate in the regulation of gene expression are poorly understood. In Escherichia coli, we have established a sequence pattern that distinguishes regulatory from nonregulatory regions. The density of promoter-like sequences, that could be recognizable by RNA polymerase and may function as potential promoters, is high within regulatory regions, in contrast to coding regions and regions located between convergently transcribed genes. Moreover, functional promoter sites identified experimentally are often found in the subregions of highest density of promoter-like signals, even when individual sites with higher binding affinity for RNA polymerase exist elsewhere within the regulatory region. In order to see the generality of this pattern, we have analyzed 43 additional genomes belonging to most established bacterial phyla. Differential densities between regulatory and nonregulatory regions are detectable in most of the analyzed genomes, with the exception of those that have evolved toward extreme genome reduction. Thus, presence of this pattern follows that of genes and other genomic features that require weak selection to be effective in order to persist. On this basis, we suggest that the loss of differential densities in the reduced genomes of host-restricted pathogens and symbionts is an outcome of the process of genome degradation resulting from the decreased efficiency of purifying selection in highly structured small populations. This implies that the differential distribution of promoter-like signals between regulatory and nonregulatory regions detected in large bacterial genomes confers a significant, although small, fitness advantage. This study paves the way for further identification of the specific types of selective constraints that affect the organization of regulatory regions and the overall distribution of promoter-like signals through more detailed comparative analyses among closely related bacterial genomes.
Conflict of interest statement
Competing interests.The authors have declared that no competing interests exist.
Figures
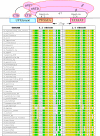

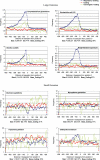
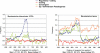
Similar articles
-
Proceedings of the SMBE Tri-National Young Investigators' Workshop 2005. Positional conservation of clusters of overlapping promoter-like sequences in enterobacterial genomes.Mol Biol Evol. 2006 May;23(5):997-1010. doi: 10.1093/molbev/msk004. Epub 2006 Mar 17. Mol Biol Evol. 2006. PMID: 16547149
-
Sigma70 promoters in Escherichia coli: specific transcription in dense regions of overlapping promoter-like signals.J Mol Biol. 2003 Oct 17;333(2):261-78. doi: 10.1016/j.jmb.2003.07.017. J Mol Biol. 2003. PMID: 14529615
-
Selection against spurious promoter motifs correlates with translational efficiency across bacteria.PLoS One. 2007 Aug 15;2(8):e745. doi: 10.1371/journal.pone.0000745. PLoS One. 2007. PMID: 17710145 Free PMC article.
-
The functional and regulatory roles of sigma factors in transcription.Cold Spring Harb Symp Quant Biol. 1998;63:141-55. doi: 10.1101/sqb.1998.63.141. Cold Spring Harb Symp Quant Biol. 1998. PMID: 10384278 Review. No abstract available.
-
Consensus architecture of promoters and transcription units in Escherichia coli: design principles for synthetic biology.Mol Biosyst. 2017 Mar 28;13(4):665-676. doi: 10.1039/c6mb00789a. Mol Biosyst. 2017. PMID: 28256660 Review.
Cited by
-
Engineering of Rhodococcus cell catalysts for tolerance improvement by sigma factor mutation and active plasmid partition.J Ind Microbiol Biotechnol. 2012 Oct;39(10):1421-30. doi: 10.1007/s10295-012-1146-5. Epub 2012 May 26. J Ind Microbiol Biotechnol. 2012. PMID: 22638790
-
Synthetic gene-regulatory networks in the opportunistic human pathogen Streptococcus pneumoniae.Proc Natl Acad Sci U S A. 2020 Nov 3;117(44):27608-27619. doi: 10.1073/pnas.1920015117. Epub 2020 Oct 21. Proc Natl Acad Sci U S A. 2020. PMID: 33087560 Free PMC article.
-
Redefining fundamental concepts of transcription initiation in bacteria.Nat Rev Genet. 2020 Nov;21(11):699-714. doi: 10.1038/s41576-020-0254-8. Epub 2020 Jul 14. Nat Rev Genet. 2020. PMID: 32665585 Free PMC article. Review.
-
RNA-Seq analysis of the multipartite genome of Rhizobium etli CE3 shows different replicon contributions under heat and saline shock.BMC Genomics. 2014 Sep 8;15(1):770. doi: 10.1186/1471-2164-15-770. BMC Genomics. 2014. PMID: 25201548 Free PMC article.
-
Mechanisms and evolution of control logic in prokaryotic transcriptional regulation.Microbiol Mol Biol Rev. 2009 Sep;73(3):481-509, Table of Contents. doi: 10.1128/MMBR.00037-08. Microbiol Mol Biol Rev. 2009. PMID: 19721087 Free PMC article. Review.
References
-
- Huerta AM, Collado-Vides J. Sigma70 promoters in Escherichia coli: Specific transcription in dense regions of overlapping promoter-like signals. J Mol Biol. 2003;333:261–278. - PubMed
-
- Gruber TM, Gross CA. Multiple sigma subunits and the partitioning of bacterial transcription space. Annu Rev Microbiol. 2003;57:441–466. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

