Murine cytomegalovirus encodes a stable intron that facilitates persistent replication in the mouse
- PMID: 17105807
- PMCID: PMC1838746
- DOI: 10.1073/pnas.0608718103
Murine cytomegalovirus encodes a stable intron that facilitates persistent replication in the mouse
Abstract
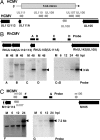
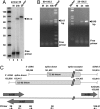
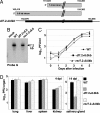
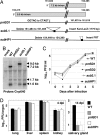
The human cytomegalovirus immediate-early 5-kb RNA previously has been shown to be a stable intron that is not required for efficient replication of the virus in cultured fibroblasts. Here we describe a murine cytomegalovirus 7.2-kb ortholog of the human cytomegalovirus 5-kb RNA. The 5' end of the 7.2-kb transcript maps to a consensus splice-donor site that is conserved among all cytomegaloviruses. We constructed mutant viruses lacking the entire 7.2-kb coding domain, the splice-donor site predicted to function in the generation of the intron or a hairpin predicted to stabilize the intron. All three mutant viruses failed to produce the 7.2-kb RNA, supporting our conclusion that it is a stable intron. Each of the mutants replicated with normal kinetics in cultured fibroblasts, but the mutants exhibited a clear defect within infected mice. Although the initial acute phase at 4 days after infection appeared to be normal, none of the mutant viruses progressed to the persistent phase, i.e., little virus was detected in the salivary gland at 14 days after infection. The intron functions as an in vivo virulence factor, facilitating progression from the acute to persistent phase of infection.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Molecular investigation of the 7.2 kb RNA of murine cytomegalovirus.Virol J. 2013 Dec 2;10:348. doi: 10.1186/1743-422X-10-348. Virol J. 2013. PMID: 24295514 Free PMC article.
-
Human cytomegalovirus 5-kilobase immediate-early RNA is a stable intron.J Virol. 2004 Dec;78(23):13182-9. doi: 10.1128/JVI.78.23.13182-13189.2004. J Virol. 2004. PMID: 15542670 Free PMC article.
-
Mutations in the temperature-sensitive murine cytomegalovirus (MCMV) mutants tsm5 and tsm30: a study of genes involved in immune evasion, DNA packaging and processing, and DNA replication.J Med Virol. 2007 Mar;79(3):285-99. doi: 10.1002/jmv.20797. J Med Virol. 2007. PMID: 17245727
-
Genetic analyses of gene function and pathogenesis of murine cytomegalovirus by transposon-mediated mutagenesis.J Clin Virol. 2002 Aug;25 Suppl 2:S111-22. doi: 10.1016/s1386-6532(02)00096-3. J Clin Virol. 2002. PMID: 12361762 Review.
-
(C)Re-combining textbook models of virus spread within the host.Cell Host Microbe. 2008 Apr 17;3(4):201-2. doi: 10.1016/j.chom.2008.03.005. Cell Host Microbe. 2008. PMID: 18407062 Review.
Cited by
-
Whole transcriptome sequencing enables discovery and analysis of viruses in archived primary central nervous system lymphomas.PLoS One. 2013 Sep 4;8(9):e73956. doi: 10.1371/journal.pone.0073956. eCollection 2013. PLoS One. 2013. PMID: 24023918 Free PMC article.
-
Characterization of Human Cytomegalovirus (HCMV) Long Non-Coding RNA1.2 During Lytic Replication.Viruses. 2025 Jan 23;17(2):149. doi: 10.3390/v17020149. Viruses. 2025. PMID: 40006904 Free PMC article.
-
Expression of pseudorabies virus-encoded long noncoding RNAs in epithelial cells and neurons.J Neurovirol. 2018 Oct;24(5):597-605. doi: 10.1007/s13365-018-0651-3. Epub 2018 Jul 9. J Neurovirol. 2018. PMID: 29987580
-
Long Non-Coding RNAs: Novel Players in Regulation of Immune Response Upon Herpesvirus Infection.Front Immunol. 2018 Apr 12;9:761. doi: 10.3389/fimmu.2018.00761. eCollection 2018. Front Immunol. 2018. PMID: 29706968 Free PMC article. Review.
-
Functional Dissection of an Alternatively Spliced Herpesvirus Gene by Splice Site Mutagenesis.J Virol. 2016 Apr 14;90(9):4626-4636. doi: 10.1128/JVI.02987-15. Print 2016 May. J Virol. 2016. PMID: 26912612 Free PMC article.
References
-
- Mocarski ES, Courcelle CT. In: Fields Virology. Knipe DM, Howley PM, editors. Vol 2. Philadelphia: Lippincott Williams & Wilkins; 2001. pp. 2629–2673.
-
- Pass RF. In: Fields Virology. Knipe DM, Howley PM, editors. Vol 2. Philadelphia: Lippincott Williams & Wilkins; 2001. pp. 2675–2706.
-
- DeMarchi JM. Virology. 1983;124:390–402. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

