Ste12 transcription factor homologue CpST12 is down-regulated by hypovirus infection and required for virulence and female fertility of the chestnut blight fungus Cryphonectria parasitica
- PMID: 17114597
- PMCID: PMC1797951
- DOI: 10.1128/EC.00302-06
Ste12 transcription factor homologue CpST12 is down-regulated by hypovirus infection and required for virulence and female fertility of the chestnut blight fungus Cryphonectria parasitica
Abstract
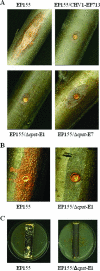
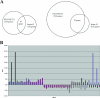
A putative homologue of the Saccharomyces cerevisiae Ste12 transcription factor was identified in a series of expressed sequence tag-based microarray analyses as being down-regulated in strains of the chestnut blight fungus, Cryphonectria parasitica, infected by virulence-attenuating hypoviruses. Cloning of the corresponding gene, cpst12, confirmed a high level of similarity to Ste12 homologues of other filamentous fungi. Disruption of cpst12 resulted in no alterations in in vitro growth characteristics or colony morphology and an increase in the production of asexual spores, indicating that CpST12 is dispensable for vegetative growth and conidiation on artificial medium. However, the disruption mutants showed a very substantial reduction in virulence on chestnut tissue and a complete loss of female fertility, two symptoms normally conferred by hypovirus infection. Both virulence and female fertility were restored by complementation with the wild-type cpst12 gene. Analysis of transcriptional changes caused by cpst12 gene disruption with a custom C. parastica cDNA microaray chip identified 152 responsive genes. A significant number of these putative CpST12-regulated genes were also responsive to hypovirus infection. Thus, cpst12 encodes a cellular transcription factor, CpST12, that is down-regulated by hypovirus infection and required for female fertility, virulence and regulated expression of a subset of hypovirus responsive host genes.
Figures


Similar articles
-
Hypovirus-responsive transcription factor gene pro1 of the chestnut blight fungus Cryphonectria parasitica is required for female fertility, asexual spore development, and stable maintenance of hypovirus infection.Eukaryot Cell. 2009 Mar;8(3):262-70. doi: 10.1128/EC.00338-08. Epub 2008 Dec 29. Eukaryot Cell. 2009. PMID: 19114501 Free PMC article.
-
Comparative analysis of alterations in host phenotype and transcript accumulation following hypovirus and mycoreovirus infections of the chestnut blight fungus Cryphonectria parasitica.Eukaryot Cell. 2007 Aug;6(8):1286-98. doi: 10.1128/EC.00166-07. Epub 2007 Jun 8. Eukaryot Cell. 2007. PMID: 17557883 Free PMC article.
-
Use of cDNA microarrays to monitor transcriptional responses of the chestnut blight fungus Cryphonectria parasitica to infection by virulence-attenuating hypoviruses.Eukaryot Cell. 2003 Dec;2(6):1253-65. doi: 10.1128/EC.2.6.1253-1265.2003. Eukaryot Cell. 2003. PMID: 14665460 Free PMC article.
-
Hypoviruses and chestnut blight: exploiting viruses to understand and modulate fungal pathogenesis.Annu Rev Genet. 2001;35:1-29. doi: 10.1146/annurev.genet.35.102401.085929. Annu Rev Genet. 2001. PMID: 11700275 Review.
-
Hypovirus molecular biology: from Koch's postulates to host self-recognition genes that restrict virus transmission.Adv Virus Res. 2013;86:109-47. doi: 10.1016/B978-0-12-394315-6.00005-2. Adv Virus Res. 2013. PMID: 23498905 Review.
Cited by
-
Deletion of the cpku80 gene in the chestnut blight fungus, Cryphonectria parasitica, enhances gene disruption efficiency.Curr Genet. 2008 Jan;53(1):59-66. doi: 10.1007/s00294-007-0162-x. Epub 2007 Oct 31. Curr Genet. 2008. PMID: 17972079
-
Cryphonectria parasitica, the causal agent of chestnut blight: invasion history, population biology and disease control.Mol Plant Pathol. 2018 Jan;19(1):7-20. doi: 10.1111/mpp.12542. Epub 2017 Apr 24. Mol Plant Pathol. 2018. PMID: 28142223 Free PMC article. Review.
-
The transcriptional regulators SteA and StuA contribute to keratin degradation and sexual reproduction of the dermatophyte Arthroderma benhamiae.Curr Genet. 2017 Feb;63(1):103-116. doi: 10.1007/s00294-016-0608-0. Epub 2016 May 11. Curr Genet. 2017. PMID: 27170358
-
Characterization of CpSte11, a MAPKKK gene of Cryphonectria parasitica, and initial evidence of its involvement in the pheromone response pathway.Mol Plant Pathol. 2012 Apr;13(3):240-50. doi: 10.1111/j.1364-3703.2011.00742.x. Epub 2011 Sep 6. Mol Plant Pathol. 2012. PMID: 21895942 Free PMC article.
-
Hypovirus-responsive transcription factor gene pro1 of the chestnut blight fungus Cryphonectria parasitica is required for female fertility, asexual spore development, and stable maintenance of hypovirus infection.Eukaryot Cell. 2009 Mar;8(3):262-70. doi: 10.1128/EC.00338-08. Epub 2008 Dec 29. Eukaryot Cell. 2009. PMID: 19114501 Free PMC article.
References
-
- Anagnostakis, S. L. 1982. Biological control of chestnut blight. Science 215:466-471. - PubMed
-
- Anagnostakis, S. L. 1984. Nuclear gene mutations in Endothia (Cryphonectria) parasitica that affect morphology and virulence. Phytopathology 74:761-765.
-
- Bowden, C. C., E. Smalley, R. P. Guries, M. Hubbes, B. Temple, and P. A. Horgen. 1996. Lack of association between cerato-ulmin production and virulence in Ophiostoma novo-ulmi. Mol. Plant-Microbe Interact. 9:556-564. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

