The mouse Gene Expression Database (GXD): 2007 update
- PMID: 17130151
- PMCID: PMC1716716
- DOI: 10.1093/nar/gkl1003
The mouse Gene Expression Database (GXD): 2007 update
Abstract
The Gene Expression Database (GXD) provides the scientific community with an extensive and easily searchable database of gene expression information about the mouse. Its primary emphasis is on developmental studies. By integrating different types of expression data, GXD aims to provide comprehensive information about expression patterns of transcripts and proteins in wild-type and mutant mice. Integration with the other Mouse Genome Informatics (MGI) databases places the gene expression information in the context of genetic, sequence, functional and phenotypic information, enabling valuable insights into the molecular biology that underlies developmental and disease processes. In recent years the utility of GXD has been greatly enhanced by a large increase in data content, obtained from the literature and provided by researchers doing large-scale in situ and cDNA screens. In addition, we have continued to refine our query and display features to make it easier for users to interrogate the data. GXD is available through the MGI web site at http://www.informatics.jax.org/ or directly at http://www.informatics.jax.org/menus/expression_menu.shtml.
Figures

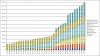

References
-
- Eppig J.T., Blake J.A., Davisson M.T., Kadin J.A., Richardson J.E., the Mouse Genome Database staff The Mouse Genome Database: a resource for today and tomorrow. Lab Anim. (NY) 2000;29:39–43. - PubMed

