Drosophila NMNAT maintains neural integrity independent of its NAD synthesis activity
- PMID: 17132048
- PMCID: PMC1665629
- DOI: 10.1371/journal.pbio.0040416
Drosophila NMNAT maintains neural integrity independent of its NAD synthesis activity
Abstract
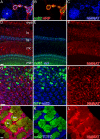
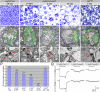
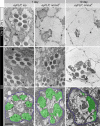
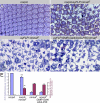
Wallerian degeneration refers to a loss of the distal part of an axon after nerve injury. Wallerian degeneration slow (Wld(s)) mice overexpress a chimeric protein containing the NAD synthase NMNAT (nicotinamide mononucleotide adenylyltransferase 1) and exhibit a delay in axonal degeneration. Currently, conflicting evidence raises questions as to whether NMNAT is the protecting factor and whether its enzymatic activity is required for such a possible function. Importantly, the link between nmnat and axon degeneration is at present solely based on overexpression studies of enzymatically active protein. Here we use the visual system of Drosophila as a model system to address these issues. We have isolated the first nmnat mutations in a multicellular organism in a forward genetic screen for synapse malfunction in Drosophila. Loss of nmnat causes a rapid and severe neurodegeneration that can be attenuated by blocking neuronal activity. Furthermore, in vivo neuronal expression of mutated nmnat shows that enzymatically inactive NMNAT protein retains strong neuroprotective effects and rescues the degeneration phenotype caused by loss of nmnat. Our data indicate an NAD-independent requirement of NMNAT for maintaining neuronal integrity that can be exploited to protect neurons from neuronal activity-induced degeneration by overexpression of the protein.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures





Comment in
-
How to protect fly photoreceptors.PLoS Biol. 2006 Dec;4(12):e438. doi: 10.1371/journal.pbio.0040438. Epub 2006 Nov 28. PLoS Biol. 2006. PMID: 20076514 Free PMC article. No abstract available.
References
-
- Zhai Q, Wang J, Kim A, Liu Q, Watts R, et al. Involvement of the ubiquitin-proteasome system in the early stages of wallerian degeneration. Neuron. 2003;39:217–225. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

