Three-dimensional models of individual cardiac histoanatomy: tools and challenges
- PMID: 17132791
- PMCID: PMC3313659
- DOI: 10.1196/annals.1380.023
Three-dimensional models of individual cardiac histoanatomy: tools and challenges
Abstract
There is a need for, and utility in, the acquisition of data sets of cardiac histoanatomy, with the vision of reconstructing individual hearts on the basis of noninvasive imaging, such as MRI, enriched by reference to detailed atlases of serial histology obtained from representative samples. These data sets would be useful not only as a repository of knowledge regarding the specifics of cardiac histoanatomy, but could form the basis for generation of individualized high-resolution cardiac structure-function models. The current article presents a step in this general direction: it illustrates how whole-heart noninvasive imaging can be combined with whole-heart histology in an approach to achieve automated construction of histoanatomically detailed models of cardiac 3D structure and function at hitherto unprecedented resolution and accuracy (based on 26.4 x 26.4 x 24.4 microm MRI voxel size, and enriched by histological detail). It provides an overview of the tools used in this quest and outlines challenges posed by the approach in the light of applications that may benefit from the availability of such data and tools.
Figures





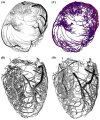


References
-
- Katz AM, Katz PB. Homogeneity out of heterogeneity. Circulation. 1989;79:712–717. - PubMed
-
- Kohl P. Cardiac cellular heterogeneity and remodelling. Cardiovasc Res. 2004;64:195–197. - PubMed
-
- Kohl P. Heterogeneous cell coupling in the heart. Circ Res. 2003;93:381–383. - PubMed
-
- Camelliti P, Green CR, LeGrice I, Kohl P. Fibroblast network in rabbit sino-atrial node: structural and functional identification of homo- and heterologous cell coupling. Circ Res. 2004;94:828–835. - PubMed
-
- Kohl P, Hunter P, Noble D. Stretch-induced changes in heart rate and rhythm: clinical observations, experiments and mathematical models. Prog Biophys Mol Biol. 1999;71:91–138. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

