BioMoby extensions to the Taverna workflow management and enactment software
- PMID: 17137515
- PMCID: PMC1693925
- DOI: 10.1186/1471-2105-7-523
BioMoby extensions to the Taverna workflow management and enactment software
Abstract
Background: As biology becomes an increasingly computational science, it is critical that we develop software tools that support not only bioinformaticians, but also bench biologists in their exploration of the vast and complex data-sets that continue to build from international genomic, proteomic, and systems-biology projects. The BioMoby interoperability system was created with the goal of facilitating the movement of data from one Web-based resource to another to fulfill the requirements of non-expert bioinformaticians. In parallel with the development of BioMoby, the European myGrid project was designing Taverna, a bioinformatics workflow design and enactment tool. Here we describe the marriage of these two projects in the form of a Taverna plug-in that provides access to many of BioMoby's features through the Taverna interface.
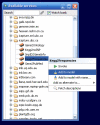
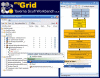
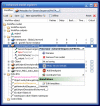
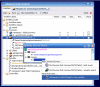
Results: The exposed BioMoby functionality aids in the design of "sensible" BioMoby workflows, aids in pipelining BioMoby and non-BioMoby-based resources, and ensures that end-users need only a minimal understanding of both BioMoby, and the Taverna interface itself. Users are guided through the construction of syntactically and semantically correct workflows through plug-in calls to the Moby Central registry. Moby Central provides a menu of only those BioMoby services capable of operating on the data-type(s) that exist at any given position in the workflow. Moreover, the plug-in automatically and correctly connects a selected service into the workflow such that users are not required to understand the nature of the inputs or outputs for any service, leaving them to focus on the biological meaning of the workflow they are constructing, rather than the technical details of how the services will interoperate.
Conclusion: With the availability of the BioMoby plug-in to Taverna, we believe that BioMoby-based Web Services are now significantly more useful and accessible to bench scientists than are more traditional Web Services.
Figures




Similar articles
-
Gbrowse Moby: a Web-based browser for BioMoby Services.Source Code Biol Med. 2006 Oct 24;1:4. doi: 10.1186/1751-0473-1-4. Source Code Biol Med. 2006. PMID: 17147784 Free PMC article.
-
Biowep: a workflow enactment portal for bioinformatics applications.BMC Bioinformatics. 2007 Mar 8;8 Suppl 1(Suppl 1):S19. doi: 10.1186/1471-2105-8-S1-S19. BMC Bioinformatics. 2007. PMID: 17430563 Free PMC article.
-
Taverna: a tool for the composition and enactment of bioinformatics workflows.Bioinformatics. 2004 Nov 22;20(17):3045-54. doi: 10.1093/bioinformatics/bth361. Epub 2004 Jun 16. Bioinformatics. 2004. PMID: 15201187
-
Interoperability with Moby 1.0--it's better than sharing your toothbrush!Brief Bioinform. 2008 May;9(3):220-31. doi: 10.1093/bib/bbn003. Epub 2008 Jan 31. Brief Bioinform. 2008. PMID: 18238804 Review.
-
Evolution of web services in bioinformatics.Brief Bioinform. 2005 Jun;6(2):178-88. doi: 10.1093/bib/6.2.178. Brief Bioinform. 2005. PMID: 15975226 Review.
Cited by
-
Update of ASRP: the Arabidopsis Small RNA Project database.Nucleic Acids Res. 2008 Jan;36(Database issue):D982-5. doi: 10.1093/nar/gkm997. Epub 2007 Nov 13. Nucleic Acids Res. 2008. PMID: 17999994 Free PMC article.
-
CoryneCenter - an online resource for the integrated analysis of corynebacterial genome and transcriptome data.BMC Syst Biol. 2007 Nov 22;1:55. doi: 10.1186/1752-0509-1-55. BMC Syst Biol. 2007. PMID: 18034885 Free PMC article.
-
Efficient, Distributed and Interactive Neuroimaging Data Analysis Using the LONI Pipeline.Front Neuroinform. 2009 Jul 20;3:22. doi: 10.3389/neuro.11.022.2009. eCollection 2009. Front Neuroinform. 2009. PMID: 19649168 Free PMC article.
-
DrugBank: a knowledgebase for drugs, drug actions and drug targets.Nucleic Acids Res. 2008 Jan;36(Database issue):D901-6. doi: 10.1093/nar/gkm958. Epub 2007 Nov 29. Nucleic Acids Res. 2008. PMID: 18048412 Free PMC article.
-
The iPlant Collaborative: Cyberinfrastructure for Plant Biology.Front Plant Sci. 2011 Jul 25;2:34. doi: 10.3389/fpls.2011.00034. eCollection 2011. Front Plant Sci. 2011. PMID: 22645531 Free PMC article.
References
-
- Wilkinson MD, Gessler DD, Farmer A, Stein L. The BioMOBY project explores open-source, simple, extensible protocols for enabling biological database interoperability. Proceedings of the Virtual Conference on Genomics and Bioinformatics. 2003;3:16–26.
-
- The BioMoby Project Homepage http://www.biomoby.org
-
- The myGrid Project Homepage http://www.mygrid.org.uk
-
- Stevens R, Robinson A, Goble CA. myGrid: Personalised Bioinformatics on the Information Grid. proceedings of 11th International Conference on Intelligent Systems in Molecular Biology, 29th June–3rd July Brisbane, Australia Bioinformatics. 2003;19:i302–i304. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

