Antigen presentation and the ubiquitin-proteasome system in host-pathogen interactions
- PMID: 17145306
- PMCID: PMC7112114
- DOI: 10.1016/S0065-2776(06)92006-9
Antigen presentation and the ubiquitin-proteasome system in host-pathogen interactions
Abstract
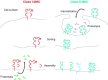
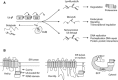
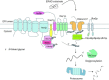
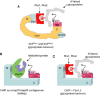
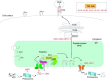
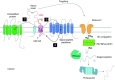
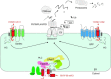
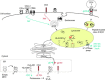
Relatively small genomes and high replication rates allow viruses and bacteria to accumulate mutations. This continuously presents the host immune system with new challenges. On the other side of the trenches, an increasingly well-adjusted host immune response, shaped by coevolutionary history, makes a pathogen's life a rather complicated endeavor. It is, therefore, no surprise that pathogens either escape detection or modulate the host immune response, often by redirecting normal cellular pathways to their advantage. For the purpose of this chapter, we focus mainly on the manipulation of the class I and class II major histocompatibility complex (MHC) antigen presentation pathways and the ubiquitin (Ub)-proteasome system by both viral and bacterial pathogens. First, we describe the general features of antigen presentation pathways and the Ub-proteasome system and then address how they are manipulated by pathogens. We discuss the many human cytomegalovirus (HCMV)-encoded immunomodulatory genes that interfere with antigen presentation (immunoevasins) and focus on the HCMV immunoevasins US2 and US11, which induce the degradation of class I MHC heavy chains by the proteasome by catalyzing their export from the endoplasmic reticulum (ER)-membrane into the cytosol, a process termed ER dislocation. US2- and US11-mediated subversion of ER dislocation ensures proteasomal degradation of class I MHC molecules and presumably allows HCMV to avoid recognition by cytotoxic T cells, whilst providing insight into general aspects of ER-associated degradation (ERAD) which is used by eukaryotic cells to purge their ER of defective proteins. We discuss the similarities and differences between the distinct pathways co-opted by US2 and US11 for dislocation and degradation of human class I MHC molecules and also a putatively distinct pathway utilized by the murine herpes virus (MHV)-68 mK3 immunoevasin for ER dislocation of murine class I MHC. We speculate on the implications of the three pathogen-exploited dislocation pathways to cellular ER quality control. Moreover, we discuss the ubiquitin (Ub)-proteasome system and its position at the core of antigen presentation as proteolysis and intracellular trafficking rely heavily on Ub-dependent processes. We add a few examples of manipulation of the Ub-proteasome system by pathogens in the context of the immune system and such diverse aspects of the host-pathogen relationship as virus budding, bacterial chromosome integration, and programmed cell death, to name a few. Finally, we speculate on newly found pathogen-encoded deubiquitinating enzymes (DUBs) and their putative roles in modulation of host-pathogen interactions.
Figures
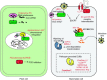
References
-
- Adams J. The proteasome: Structure, function, and role in the cell. Cancer Treat. Rev. 2003;29(Suppl. 1):3–9. - PubMed
-
- Ahn K., Gruhler A., Galocha B., Jones T.R., Wiertz E.J., Ploegh H.L., Peterson P.A., Yang Y., Fruh K. The ER‐luminal domain of the HCMV glycoprotein US6 inhibits peptide translocation by TAP. Immunity. 1997;6(5):613–621. - PubMed
-
- Aiken C., Konner J., Landau N.R., Lenburg M.E., Trono D. Nef induces CD4 endocytosis: Requirement for a critical dileucine motif in the membrane‐proximal CD4 cytoplasmic domain. Cell. 1994;76(5):853–864. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

