Molecular basis of Ca(v)2.3 calcium channels in rat nociceptive neurons
- PMID: 17145762
- PMCID: PMC2760738
- DOI: 10.1074/jbc.M605248200
Molecular basis of Ca(v)2.3 calcium channels in rat nociceptive neurons
Abstract
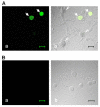
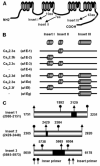
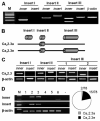
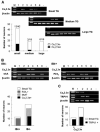
Ca(v)2.3 calcium channels play an important role in pain transmission in peripheral sensory neurons. Six Ca(v)2.3 isoforms resulting from different combinations of three inserts (inserts I and II in the II-III loop and insert III in the carboxyl-terminal region) have been identified in different mammalian tissues. To date, however, Ca(v)2.3 isoforms unique to primary sensory neurons have not been identified. In this study, we determined Ca(v)2.3 isoforms expressed in the rat trigeminal ganglion neurons. Whole tissue reverse transcription (RT)-PCR analyses revealed that only two isoforms, Ca(v)2.3a and Ca(v)2.3e, are present in TG neurons. Using single cell RT-PCR, we found that Ca(v)2.3e is the major isoform, whereas Ca(v)2.3e expression is highly restricted to small (<16 mum) isolectin B4-negative and tyrosine kinase A-positive neurons. Ca(v)2.3e was also preferentially detected in neurons expressing the nociceptive marker, transient receptor potential vanilloid 1. Single cell RT-PCR following calcium imaging and whole-cell patch clamp recordings provided evidence of an association between an R-type calcium channel component and Ca(v)2.3e expression. Our results suggest that Ca(v)2.3e in sensory neurons may be a potential target for the treatment of pain.
Figures

Similar articles
-
R-type Calcium Channel Isoform in Rat Dorsal Root Ganglion Neurons.Korean J Physiol Pharmacol. 2010 Feb;14(1):45-9. doi: 10.4196/kjpp.2010.14.1.45. Epub 2010 Feb 28. Korean J Physiol Pharmacol. 2010. PMID: 20221279 Free PMC article.
-
Relative roles of protein kinase A and protein kinase C in modulation of transient receptor potential vanilloid type 1 receptor responsiveness in rat sensory neurons in vitro and peripheral nociceptors in vivo.Neuroscience. 2006 Jun 30;140(2):645-57. doi: 10.1016/j.neuroscience.2006.02.035. Epub 2006 Mar 29. Neuroscience. 2006. PMID: 16564637
-
Signaling mechanisms of down-regulation of voltage-activated Ca2+ channels by transient receptor potential vanilloid type 1 stimulation with olvanil in primary sensory neurons.Neuroscience. 2006 Aug 11;141(1):407-19. doi: 10.1016/j.neuroscience.2006.03.023. Epub 2006 May 6. Neuroscience. 2006. PMID: 16678970
-
Transient receptor potential V2 expressed in sensory neurons is activated by probenecid.Neurosci Lett. 2007 Sep 25;425(2):120-5. doi: 10.1016/j.neulet.2007.08.035. Epub 2007 Aug 24. Neurosci Lett. 2007. PMID: 17850966
-
Targeting Nociceptive Neurons and Transient Receptor Potential Channels for the Treatment of Migraine.Int J Mol Sci. 2023 Apr 26;24(9):7897. doi: 10.3390/ijms24097897. Int J Mol Sci. 2023. PMID: 37175602 Free PMC article. Review.
Cited by
-
Effects of familial hemiplegic migraine type 1 mutation T666M on voltage-gated calcium channel activities in trigeminal ganglion neurons.J Neurophysiol. 2012 Mar;107(6):1666-80. doi: 10.1152/jn.00551.2011. Epub 2011 Dec 21. J Neurophysiol. 2012. PMID: 22190617 Free PMC article.
-
CRMP2 and voltage-gated ion channels: potential roles in neuropathic pain.Neuronal Signal. 2018;2(1):NS20170220. doi: 10.1042/NS20170220. Epub 2018 Mar 30. Neuronal Signal. 2018. PMID: 30364788 Free PMC article.
-
How "Pharmacoresistant" is Cav2.3, the Major Component of Voltage-Gated R-type Ca2+ Channels?Pharmaceuticals (Basel). 2013 May 27;6(6):759-76. doi: 10.3390/ph6060759. Pharmaceuticals (Basel). 2013. PMID: 24276260 Free PMC article.
-
Effect of amitriptyline on tetrodotoxin-resistant Nav1.9 currents in nociceptive trigeminal neurons.Mol Pain. 2013 Jun 22;9:31. doi: 10.1186/1744-8069-9-31. Mol Pain. 2013. PMID: 24228717 Free PMC article.
-
TLR4 enhances histamine-mediated pruritus by potentiating TRPV1 activity.Mol Brain. 2014 Aug 21;7:59. doi: 10.1186/s13041-014-0059-9. Mol Brain. 2014. PMID: 25139109 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

