Almost all human genes resulted from ancient duplication
- PMID: 17146051
- PMCID: PMC1748171
- DOI: 10.1073/pnas.0608796103
Almost all human genes resulted from ancient duplication
Abstract
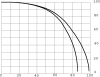
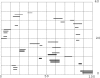
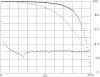
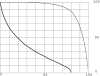
Results of protein sequence comparison at open criterion show a very large number of relationships that have, up to now, gone unreported. The relationships suggest many ancient events of gene duplication. It is well known that gene duplication has been a major process in the evolution of genomes. A collection of human genes that have known functions have been examined for a history of gene duplications detected by means of amino acid sequence similarity by using BLASTp with an expectation of two or less (open criterion). Because the collection of genes in build 35 includes sets of transcript variants, all genes of known function were collected, and only the longest transcription variant was included, yielding a 13,298-member library called KGMV (for known genes maximum variant). When all lengths of matches are accepted, >97% of human genes show significant matches to each other. Many form matches with a large number of other different proteins, showing that most genes are made up from parts of many others as a result of ancient events of duplication. To support the use of the open criterion, all of the members of the KGMV library were twice replaced with random protein sequences of the same length and average composition, and all were compared with each other with BLASTp at expectation two or less. The set of matches averaged 0.35% of that observed for the KGMV set of proteins.
Conflict of interest statement
The author declares no conflict of interest.
Figures

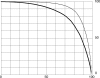
Similar articles
-
Transposable elements have contributed to thousands of human proteins.Proc Natl Acad Sci U S A. 2006 Feb 7;103(6):1798-803. doi: 10.1073/pnas.0510007103. Epub 2006 Jan 27. Proc Natl Acad Sci U S A. 2006. PMID: 16443682 Free PMC article.
-
The majority of human genes have regions repeated in other human genes.Proc Natl Acad Sci U S A. 2005 Apr 12;102(15):5466-70. doi: 10.1073/pnas.0501008102. Epub 2005 Mar 31. Proc Natl Acad Sci U S A. 2005. PMID: 15802472 Free PMC article.
-
Adaptive divergence of ancient gene duplicates in the avian MHC class II beta.Mol Biol Evol. 2010 Oct;27(10):2360-74. doi: 10.1093/molbev/msq120. Epub 2010 May 12. Mol Biol Evol. 2010. PMID: 20463048
-
Evolution of duplications in the transferrin family of proteins.Comp Biochem Physiol B Biochem Mol Biol. 2005 Jan;140(1):11-25. doi: 10.1016/j.cbpc.2004.09.012. Comp Biochem Physiol B Biochem Mol Biol. 2005. PMID: 15621505 Review.
-
Computational approaches to unveiling ancient genome duplications.Nat Rev Genet. 2004 Oct;5(10):752-63. doi: 10.1038/nrg1449. Nat Rev Genet. 2004. PMID: 15510166 Review.
Cited by
-
Pervasive and persistent redundancy among duplicated genes in yeast.PLoS Genet. 2008 Jul 4;4(7):e1000113. doi: 10.1371/journal.pgen.1000113. PLoS Genet. 2008. PMID: 18604285 Free PMC article.
-
The evolutionary dynamics of the Saccharomyces cerevisiae protein interaction network after duplication.Proc Natl Acad Sci U S A. 2008 Jan 22;105(3):950-4. doi: 10.1073/pnas.0707293105. Epub 2008 Jan 16. Proc Natl Acad Sci U S A. 2008. PMID: 18199840 Free PMC article.
-
The role of duplications in the evolution of genomes highlights the need for evolutionary-based approaches in comparative genomics.Biol Direct. 2011 Feb 18;6:11. doi: 10.1186/1745-6150-6-11. Biol Direct. 2011. PMID: 21333002 Free PMC article. Review.
-
Phylogenetic and functional characterization of the hAT transposon superfamily.Genetics. 2011 May;188(1):45-57. doi: 10.1534/genetics.111.126813. Epub 2011 Mar 2. Genetics. 2011. PMID: 21368277 Free PMC article.
-
An Overview of Duplicated Gene Detection Methods: Why the Duplication Mechanism Has to Be Accounted for in Their Choice.Genes (Basel). 2020 Sep 4;11(9):1046. doi: 10.3390/genes11091046. Genes (Basel). 2020. PMID: 32899740 Free PMC article. Review.
References
-
- Ohno S. Evolution by Gene Duplication. New York: Springer; 1970.
-
- Britten RJ. Carnegie Institution Yearbook 64. Washington, DC: Carnegie Institution; 1965. p. 333.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

