Computational analyses of the surface properties of protein-protein interfaces
- PMID: 17164526
- PMCID: PMC2483497
- DOI: 10.1107/S0907444906046762
Computational analyses of the surface properties of protein-protein interfaces
Abstract
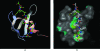
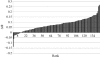
Several potential applications of structural biology depend on discovering how one macromolecule might recognize a partner. Experiment remains the best way to answer this question, but computational tools can contribute where this fails. In such cases, structures may be studied to identify patches of exposed residues that have properties common to interaction surfaces and the locations of these patches can serve as the basis for further modelling or for further experimentation. To date, interaction surfaces have been proposed on the basis of unusual physical properties, unusual propensities for particular amino-acid types or an unusually high level of sequence conservation. Using the CXXSurface toolkit, developed as a part of the CCP4MG program, a suite of tools to analyse the properties of surfaces and their interfaces in complexes has been prepared and applied. These tools have enabled the rapid analysis of known complexes to evaluate the distribution of (i) hydrophobicity, (ii) electrostatic complementarity and (iii) sequence conservation in authentic complexes, so as to assess the extent to which these properties may be useful indicators of probable biological function.
Figures



Similar articles
-
Predicting protein interaction sites: binding hot-spots in protein-protein and protein-ligand interfaces.Bioinformatics. 2006 Jun 1;22(11):1335-42. doi: 10.1093/bioinformatics/btl079. Epub 2006 Mar 7. Bioinformatics. 2006. PMID: 16522669
-
Protein-protein docking using 3D-Dock in rounds 3, 4, and 5 of CAPRI.Proteins. 2005 Aug 1;60(2):281-8. doi: 10.1002/prot.20571. Proteins. 2005. PMID: 15981271
-
Scoring docking models with evolutionary information.Proteins. 2005 Aug 1;60(2):275-80. doi: 10.1002/prot.20570. Proteins. 2005. PMID: 15981273
-
The "Sticky Patch" Model of Crystallization and Modification of Proteins for Enhanced Crystallizability.Methods Mol Biol. 2017;1607:77-115. doi: 10.1007/978-1-4939-7000-1_4. Methods Mol Biol. 2017. PMID: 28573570 Free PMC article. Review.
-
Electrostatic aspects of protein-protein interactions.Curr Opin Struct Biol. 2000 Apr;10(2):153-9. doi: 10.1016/s0959-440x(00)00065-8. Curr Opin Struct Biol. 2000. PMID: 10753808 Review.
Cited by
-
In-silico analysis of the strigolactone ligand-receptor system.Plant Direct. 2020 Sep 15;4(9):e00263. doi: 10.1002/pld3.263. eCollection 2020 Sep. Plant Direct. 2020. PMID: 32995702 Free PMC article.
-
Predicting where small molecules bind at protein-protein interfaces.PLoS One. 2013;8(3):e58583. doi: 10.1371/journal.pone.0058583. Epub 2013 Mar 7. PLoS One. 2013. PMID: 23505538 Free PMC article.
-
Trapping and Driving Individual Charged Micro-particles in Fluid with an Electrostatic Device.Nanomicro Lett. 2016;8(3):270-281. doi: 10.1007/s40820-016-0087-3. Epub 2016 Mar 10. Nanomicro Lett. 2016. PMID: 30460287 Free PMC article.
-
A simple procedure for the derivation of electron density based surfaces of drug-receptor complexes from a combination of X-ray data and theoretical calculations.Bioorg Med Chem. 2010 Aug 15;18(16):5965-74. doi: 10.1016/j.bmc.2010.06.080. Epub 2010 Jul 1. Bioorg Med Chem. 2010. PMID: 20634077 Free PMC article.
-
Beauty is in the eye of the beholder: proteins can recognize binding sites of homologous proteins in more than one way.PLoS Comput Biol. 2010 Jun 17;6(6):e1000821. doi: 10.1371/journal.pcbi.1000821. PLoS Comput Biol. 2010. PMID: 20585553 Free PMC article.
References
-
- Bartlett, G. J., Porter, C. T., Borkakoti, N. & Thornton, J. M. (2002). J. Mol. Biol.324, 105–121. - PubMed
-
- Brown, N. R., Noble, M. E., Endicott, J. A. & Johnson, L. N. (1999). Nature Cell Biol.1, 438–443. - PubMed
-
- Dill, K. A. (1990). Biochemistry, 29, 7133–7155. - PubMed
-
- Frank, H. & Evans, M. (1945). J. Chem. Phys.13, 507–532.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

