The coactivator function of Arabidopsis NPR1 requires the core of its BTB/POZ domain and the oxidation of C-terminal cysteines
- PMID: 17172357
- PMCID: PMC1785396
- DOI: 10.1105/tpc.106.046953
The coactivator function of Arabidopsis NPR1 requires the core of its BTB/POZ domain and the oxidation of C-terminal cysteines
Abstract
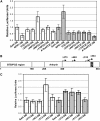
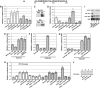
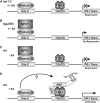
NONEXPRESSOR OF PATHOGENESIS-RELATED GENES1 (NPR1) regulates systemic acquired resistance (SAR) in Arabidopsis thaliana, and current models propose that after treatment with salicylic acid (SA), Cys-82 and Cys-216 of NPR1 are reduced, leading to nuclear import. The interaction of nucleus-localized NPR1 with TGA transcription factors results in the activation of defense genes, including the SAR marker PATHOGENESIS-RELATED-1 (PR-1), and the deployment of SAR. Little is known about how TGA factors or NPR1 regulate transcription or whether a TGA-NPR1 complex forms on DNA. We show that TGA2 and NPR1 are recruited to PR-1 independently of each other and of SA treatment. Consistent with the result that a triple knockout in TGA2/5/6 derepresses PR-1, in vivo plant transcription assays revealed that TGA2 is not an autonomous transcription activator but is a transcriptional repressor in both untreated and SA-treated cells. However, after stimulation with SA, TGA2 is incorporated into a transactivating complex with NPR1, forming an enhanceosome that requires the core of the NPR1 BTB/POZ domain (residues 80 to 91) and the oxidation of NPR1 Cys-521 and Cys-529. These Cys residues are found in a new type of transactivation domain that we term Cys-oxidized. These data further our understanding of the mechanism by which TGA2 and NPR1 activate Arabidopsis PR-1.
Figures


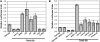
References
-
- Abate, C., Patel, L., Rauscher, F.J., III, and Curran, T. (1990). Redox regulation of fos and jun DNA-binding activity in vitro. Science 249 1157–1161. - PubMed
-
- Aravind, L., and Koonin, E.V. (1999). Fold prediction and evolutionary analysis of the POZ domain: Structural and evolutionary relationship with the potassium channel tetramerization domain. J. Mol. Biol. 285 1353–1361. - PubMed
-
- Bardwell, V.J., and Treisman, R. (1994). The POZ domain: A conserved protein-protein interaction motif. Genes Dev. 8 1664–1677. - PubMed
-
- Benezra, R. (1994). An intermolecular disulfide bond stabilizes E2A homodimers and is required for DNA binding at physiological temperatures. Cell 79 1057–1067. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

