Three-dimensional morphology and gene expression in the Drosophila blastoderm at cellular resolution I: data acquisition pipeline
- PMID: 17184546
- PMCID: PMC1794436
- DOI: 10.1186/gb-2006-7-12-r123
Three-dimensional morphology and gene expression in the Drosophila blastoderm at cellular resolution I: data acquisition pipeline
Abstract
Background: To model and thoroughly understand animal transcription networks, it is essential to derive accurate spatial and temporal descriptions of developing gene expression patterns with cellular resolution.
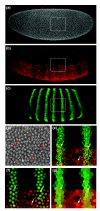
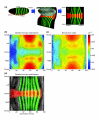
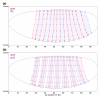
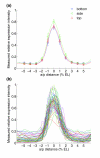
Results: Here we describe a suite of methods that provide the first quantitative three-dimensional description of gene expression and morphology at cellular resolution in whole embryos. A database containing information derived from 1,282 embryos is released that describes the mRNA expression of 22 genes at multiple time points in the Drosophila blastoderm. We demonstrate that our methods are sufficiently accurate to detect previously undescribed features of morphology and gene expression. The cellular blastoderm is shown to have an intricate morphology of nuclear density patterns and apical/basal displacements that correlate with later well-known morphological features. Pair rule gene expression stripes, generally considered to specify patterning only along the anterior/posterior body axis, are shown to have complex changes in stripe location, stripe curvature, and expression level along the dorsal/ventral axis. Pair rule genes are also found to not always maintain the same register to each other.
Conclusion: The application of these quantitative methods to other developmental systems will likely reveal many other previously unknown features and provide a more rigorous understanding of developmental regulatory networks.
Figures








Similar articles
-
Three-dimensional morphology and gene expression in the Drosophila blastoderm at cellular resolution II: dynamics.Genome Biol. 2006;7(12):R124. doi: 10.1186/gb-2006-7-12-r124. Genome Biol. 2006. PMID: 17184547 Free PMC article.
-
Three-dimensional morphology and gene expression mapping for the Drosophila blastoderm.Cold Spring Harb Protoc. 2012 Feb 1;2012(2):150-61. doi: 10.1101/pdb.top067843. Cold Spring Harb Protoc. 2012. PMID: 22301653 Free PMC article.
-
A two-step mode of stripe formation in the Drosophila blastoderm requires interactions among primary pair rule genes.Mech Dev. 1994 Jan;45(1):3-13. doi: 10.1016/0925-4773(94)90049-3. Mech Dev. 1994. PMID: 8186146
-
Drosophila blastoderm patterning.Curr Opin Genet Dev. 2012 Dec;22(6):533-41. doi: 10.1016/j.gde.2012.10.005. Epub 2013 Jan 4. Curr Opin Genet Dev. 2012. PMID: 23290311 Review.
-
Building quantitative, three-dimensional atlases of gene expression and morphology at cellular resolution.Wiley Interdiscip Rev Dev Biol. 2013 Nov-Dec;2(6):767-79. doi: 10.1002/wdev.107. Epub 2013 Feb 4. Wiley Interdiscip Rev Dev Biol. 2013. PMID: 24123936 Free PMC article. Review.
Cited by
-
Medium-throughput processing of whole mount in situ hybridisation experiments into gene expression domains.PLoS One. 2012;7(9):e46658. doi: 10.1371/journal.pone.0046658. Epub 2012 Sep 28. PLoS One. 2012. PMID: 23029561 Free PMC article.
-
Interactive processing and visualization of image data for biomedical and life science applications.BMC Cell Biol. 2007 Jul 10;8 Suppl 1(Suppl 1):S10. doi: 10.1186/1471-2121-8-S1-S10. BMC Cell Biol. 2007. PMID: 17634091 Free PMC article.
-
The gap gene network.Cell Mol Life Sci. 2011 Jan;68(2):243-74. doi: 10.1007/s00018-010-0536-y. Epub 2010 Oct 8. Cell Mol Life Sci. 2011. PMID: 20927566 Free PMC article. Review.
-
Multicolor fluorescent in situ hybridization to define abutting and overlapping gene expression in the embryonic zebrafish brain.Neural Dev. 2011 Apr 5;6:10. doi: 10.1186/1749-8104-6-10. Neural Dev. 2011. PMID: 21466670 Free PMC article.
-
Unified bursting strategies in ectopic and endogenous even-skipped expression patterns.bioRxiv [Preprint]. 2024 Jun 25:2023.02.09.527927. doi: 10.1101/2023.02.09.527927. bioRxiv. 2024. Update in: Elife. 2024 Dec 09;12:RP88671. doi: 10.7554/eLife.88671. PMID: 36798351 Free PMC article. Updated. Preprint.
References
-
- Burne RM, Bard JB, Dubreuil C, Guest E, Hill W, Kaufman M, Stark M, Davidson D, Baldock RA. A three-dimensional model of the mouse at embryonic day 9. Dev Biol. 1999;216:457–468. - PubMed
-
- Megason SG, Fraser SE. Digitizing life at the level of the cell: high-performance laser-scanning microscopy and image analysis for in toto imaging of development. Mechanisms Dev. 2003;120:1407–1420. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

