High performance computing in biology: multimillion atom simulations of nanoscale systems
- PMID: 17187988
- PMCID: PMC1868470
- DOI: 10.1016/j.jsb.2006.10.023
High performance computing in biology: multimillion atom simulations of nanoscale systems
Abstract
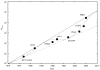
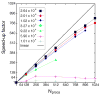
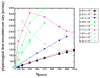
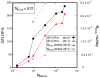
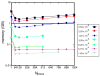
Computational methods have been used in biology for sequence analysis (bioinformatics), all-atom simulation (molecular dynamics and quantum calculations), and more recently for modeling biological networks (systems biology). Of these three techniques, all-atom simulation is currently the most computationally demanding, in terms of compute load, communication speed, and memory load. Breakthroughs in electrostatic force calculation and dynamic load balancing have enabled molecular dynamics simulations of large biomolecular complexes. Here, we report simulation results for the ribosome, using approximately 2.64 million atoms, the largest all-atom biomolecular simulation published to date. Several other nano-scale systems with different numbers of atoms were studied to measure the performance of the NAMD molecular dynamics simulation program on the Los Alamos National Laboratory Q Machine. We demonstrate that multimillion atom systems represent a 'sweet spot' for the NAMD code on large supercomputers. NAMD displays an unprecedented 85% parallel scaling efficiency for the ribosome system on 1024 CPUs. We also review recent targeted molecular dynamics simulations of the ribosome that prove useful for studying conformational changes of this large biomolecular complex in atomic detail.
Figures




References
-
- Auffinger P, Westhof E. Simulations of the molecular dynamics of nucleic acids. Current Opinion in Structural Biology. 1998;8(2):227–236. - PubMed
-
- Auffinger P, Westhof E. Water and ion binding around r(UpA)(12) and d(TpA)(12) oligomers: Comparison with RNA and DNA (CpG)(12) duplexes. Journal of Molecular Biology. 2001;305(5):1057–1072. - PubMed
-
- Auffinger P, Westhof E. Melting of the solvent structure around a RNA duplex: a molecular dynamics simulation study. Biophys Chem. 2002;95(3):203–210. - PubMed
-
- Board J, Causey J, Leathrum J, Windemuth A, Schulten K. Accelerated molecular dynamics simulation with the parallel fast multipole algorithm. Chemical Physics Letters. 1992;198:89–94.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

