A gene expression signature predicts survival of patients with stage I non-small cell lung cancer
- PMID: 17194181
- PMCID: PMC1716187
- DOI: 10.1371/journal.pmed.0030467
A gene expression signature predicts survival of patients with stage I non-small cell lung cancer
Abstract
Background: Lung cancer is the leading cause of cancer-related death in the United States. Nearly 50% of patients with stages I and II non-small cell lung cancer (NSCLC) will die from recurrent disease despite surgical resection. No reliable clinical or molecular predictors are currently available for identifying those at high risk for developing recurrent disease. As a consequence, it is not possible to select those high-risk patients for more aggressive therapies and assign less aggressive treatments to patients at low risk for recurrence.
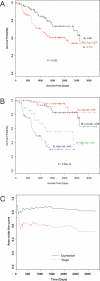
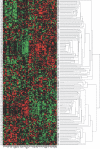
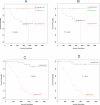
Methods and findings: In this study, we applied a meta-analysis of datasets from seven different microarray studies on NSCLC for differentially expressed genes related to survival time (under 2 y and over 5 y). A consensus set of 4,905 genes from these studies was selected, and systematic bias adjustment in the datasets was performed by distance-weighted discrimination (DWD). We identified a gene expression signature consisting of 64 genes that is highly predictive of which stage I lung cancer patients may benefit from more aggressive therapy. Kaplan-Meier analysis of the overall survival of stage I NSCLC patients with the 64-gene expression signature demonstrated that the high- and low-risk groups are significantly different in their overall survival. Of the 64 genes, 11 are related to cancer metastasis (APC, CDH8, IL8RB, LY6D, PCDHGA12, DSP, NID, ENPP2, CCR2, CASP8, and CASP10) and eight are involved in apoptosis (CASP8, CASP10, PIK3R1, BCL2, SON, INHA, PSEN1, and BIK).
Conclusions: Our results indicate that gene expression signatures from several datasets can be reconciled. The resulting signature is useful in predicting survival of stage I NSCLC and might be useful in informing treatment decisions.
Conflict of interest statement
Figures

Similar articles
-
A five-gene signature and clinical outcome in non-small-cell lung cancer.N Engl J Med. 2007 Jan 4;356(1):11-20. doi: 10.1056/NEJMoa060096. N Engl J Med. 2007. PMID: 17202451
-
Non-overlapping and non-cell-type-specific gene expression signatures predict lung cancer survival.J Clin Oncol. 2008 Feb 20;26(6):877-83. doi: 10.1200/JCO.2007.13.1516. J Clin Oncol. 2008. PMID: 18281660
-
Prediction of recurrence-free survival in postoperative non-small cell lung cancer patients by using an integrated model of clinical information and gene expression.Clin Cancer Res. 2008 Nov 15;14(22):7397-404. doi: 10.1158/1078-0432.CCR-07-4937. Clin Cancer Res. 2008. PMID: 19010856
-
Gene expression profiling for early-stage NSCLC.Am J Clin Oncol. 2015 Feb;38(1):103-7. doi: 10.1097/COC.0b013e31828d95d8. Am J Clin Oncol. 2015. PMID: 23608827 Review.
-
Aspects of lung cancer gene expression profiling.Curr Opin Drug Discov Devel. 2004 May;7(3):290-303. Curr Opin Drug Discov Devel. 2004. PMID: 15216932 Review.
Cited by
-
Identification of molecular biomarkers and pathways of NSCLC: insights from a systems biomedicine perspective.J Genet Eng Biotechnol. 2021 Mar 19;19(1):43. doi: 10.1186/s43141-021-00134-1. J Genet Eng Biotechnol. 2021. PMID: 33742334 Free PMC article.
-
Identification and validation of hypoxia-derived gene signatures to predict clinical outcomes and therapeutic responses in stage I lung adenocarcinoma patients.Theranostics. 2021 Mar 5;11(10):5061-5076. doi: 10.7150/thno.56202. eCollection 2021. Theranostics. 2021. PMID: 33754044 Free PMC article.
-
A Prognostic Model Based on Six Metabolism-Related Genes in Colorectal Cancer.Biomed Res Int. 2020 Aug 31;2020:5974350. doi: 10.1155/2020/5974350. eCollection 2020. Biomed Res Int. 2020. PMID: 32953885 Free PMC article.
-
New molecularly targeted therapies for lung cancer.J Clin Invest. 2007 Oct;117(10):2740-50. doi: 10.1172/JCI31809. J Clin Invest. 2007. PMID: 17909619 Free PMC article. Review.
-
A white-box approach to microarray probe response characterization: the BaFL pipeline.BMC Bioinformatics. 2009 Dec 29;10:449. doi: 10.1186/1471-2105-10-449. BMC Bioinformatics. 2009. PMID: 20040098 Free PMC article.
References
-
- Spiro SG, Silvestri GA. One hundred years of lung cancer. Am J Respir Crit Care Med. 2005;172:523–529. - PubMed
-
- Bach PB, Kelley MJ, Tate RC, McCrory DC. Screening for lung cancer: A review of the current literature. Chest. 2003;123:72S–82S. - PubMed
-
- Mountain CF, Dresler CM. Regional lymph node classification for lung cancer staging. Chest. 1997;111:1718–1723. - PubMed
-
- El-Sherif A, Luketich JD, Landreneau RJ, Fernando HC. New therapeutic approaches for early stage non-small cell lung cancer. Surg Oncol. 2005;14:27–32. - PubMed
-
- Beer DG, Kardia SL, Huang CC, Giordano TJ, Levin AM, et al. Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nat Med. 2002;8:816–824. - PubMed
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

