Type III effector diversification via both pathoadaptation and horizontal transfer in response to a coevolutionary arms race
- PMID: 17194219
- PMCID: PMC1713259
- DOI: 10.1371/journal.pgen.0020209
Type III effector diversification via both pathoadaptation and horizontal transfer in response to a coevolutionary arms race
Abstract
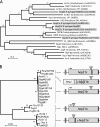
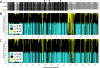
The concept of the coevolutionary arms race holds a central position in our understanding of pathogen-host interactions. Here we identify the molecular mechanisms and follow the stepwise progression of an arms race in a natural system. We show how the evolution and function of the HopZ family of type III secreted effector proteins carried by the plant pathogen Pseudomonas syringae are influenced by a coevolutionary arms race between pathogen and host. We surveyed 96 isolates of P. syringae and identified three homologs (HopZ1, HopZ2, and HopZ3) distributed among approximately 45% of the strains. All alleles were sequenced and their expression was confirmed. Evolutionary analyses determined that the diverse HopZ1 homologs are ancestral to P. syringae, and have diverged via pathoadaptive mutational changes into three functional and two degenerate forms, while HopZ2 and HopZ3 have been brought into P. syringae via horizontal transfer from other ecologically similar bacteria. A PAML selection analysis revealed that the C terminus of HopZ1 is under strong positive selection. Despite the extensive genetic variation observed in this family, all three homologs have cysteine-protease activity, although their substrate specificity may vary. The introduction of the ancestral hopZ1 allele into strains harboring alternate alleles results in a resistance protein-mediated defense response in their respective hosts, which is not observed with the endogenous allele. These data indicate that the P. syringae HopZ family has undergone allelic diversification via both pathoadaptive mutational changes and horizontal transfer in response to selection imposed by the host defense system. This genetic diversity permits the pathogen to avoid host defenses while still maintaining a virulence-associated protease, thereby allowing it to thrive on its current host, while simultaneously impacting its host range.
Conflict of interest statement
Competing interests. The authors have declared that no competing interests exist.
Figures


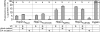


Similar articles
-
Allelic variants of the Pseudomonas syringae type III effector HopZ1 are differentially recognized by plant resistance systems.Mol Plant Microbe Interact. 2009 Feb;22(2):176-89. doi: 10.1094/MPMI-22-2-0176. Mol Plant Microbe Interact. 2009. PMID: 19132870
-
Catalytic domain of the diversified Pseudomonas syringae type III effector HopZ1 determines the allelic specificity in plant hosts.Mol Microbiol. 2010 Apr;76(2):437-55. doi: 10.1111/j.1365-2958.2010.07107.x. Epub 2010 Mar 10. Mol Microbiol. 2010. PMID: 20233307
-
Diverse evolutionary mechanisms shape the type III effector virulence factor repertoire in the plant pathogen Pseudomonas syringae.Genetics. 2004 Jul;167(3):1341-60. doi: 10.1534/genetics.103.019638. Genetics. 2004. PMID: 15280247 Free PMC article.
-
Closing the circle on the discovery of genes encoding Hrp regulon members and type III secretion system effectors in the genomes of three model Pseudomonas syringae strains.Mol Plant Microbe Interact. 2006 Nov;19(11):1151-8. doi: 10.1094/MPMI-19-1151. Mol Plant Microbe Interact. 2006. PMID: 17073298 Review.
-
Pseudomonas syringae: what it takes to be a pathogen.Nat Rev Microbiol. 2018 May;16(5):316-328. doi: 10.1038/nrmicro.2018.17. Epub 2018 Feb 26. Nat Rev Microbiol. 2018. PMID: 29479077 Free PMC article. Review.
Cited by
-
Auto-acetylation on K289 is not essential for HopZ1a-mediated plant defense suppression.Front Microbiol. 2015 Jul 8;6:684. doi: 10.3389/fmicb.2015.00684. eCollection 2015. Front Microbiol. 2015. PMID: 26217317 Free PMC article.
-
Comparative genomics reveals genes significantly associated with woody hosts in the plant pathogen Pseudomonas syringae.Mol Plant Pathol. 2016 Dec;17(9):1409-1424. doi: 10.1111/mpp.12423. Epub 2016 Jul 15. Mol Plant Pathol. 2016. PMID: 27145446 Free PMC article.
-
Comparative Genomics and Gene Pool Analysis Reveal the Decrease of Genome Diversity and Gene Number in Rice Blast Fungi by Stable Adaption with Rice.J Fungi (Basel). 2021 Dec 22;8(1):5. doi: 10.3390/jof8010005. J Fungi (Basel). 2021. PMID: 35049945 Free PMC article.
-
Membrane vesicle engineering with "à la carte" bacterial-immunogenic molecules for organism-free plant vaccination.Microb Biotechnol. 2023 Dec;16(12):2223-2235. doi: 10.1111/1751-7915.14323. Epub 2023 Aug 2. Microb Biotechnol. 2023. PMID: 37530752 Free PMC article. Review.
-
Identification of new type III effectors and analysis of the plant response by competitive index.Mol Plant Pathol. 2009 Jan;10(1):69-80. doi: 10.1111/j.1364-3703.2008.00511.x. Mol Plant Pathol. 2009. PMID: 19161354 Free PMC article.
References
-
- Dawkins R, Krebs JR. Arms races between and within species. Proc R Soc Lond B Biol Sci. 1979;205:489–511. - PubMed
-
- Bergelson J, Dwyer G, Emerson JJ. Models and data on plant-enemy coevolution. Annu Rev Genet. 2001;35:469–499. - PubMed
-
- Clay K, Kover PX. The Red Queen hypothesis and plant/pathogen interactions. Annu Rev Phytopathol. 1996;34:29–50. - PubMed
-
- Mota LJ, Cornelis GR. The bacterial injection kit: Type III secretion systems. Ann Med. 2005;37:234–249. - PubMed
-
- Jackson RW, Athanassopoulos E, Tsiamis G, Mansfield JW, Sesma A, et al. Identification of a pathogenicity island, which contains genes for virulence and avirulence, on a large native plasmid in the bean pathogen Pseudomonas syringae pathovar phaseolicola. Proc Natl Acad Sci U S A. 1999;96:10875–10880. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

