Evidence for U-tail stabilization of gRNA/mRNA interactions in kinetoplastid RNA editing
- PMID: 17194935
- PMCID: PMC2762388
- DOI: 10.4161/rna.1.1.898
Evidence for U-tail stabilization of gRNA/mRNA interactions in kinetoplastid RNA editing
Abstract
The most dramatic example of RNA editing is found in the mitochondria of trypanosomes. In these organisms, U-insertions/deletions can create mRNAs that are twice as large as the gene that encodes them. Guide RNAs (gRNAs) that are complementary to short stretches of the mature message direct the precise placements of the U residues. The binding of gRNA to mRNA is a fundamental step in RNA editing and understanding the relative importance of the elements that confer affinity and specificity on this interaction is critical to our understanding of the editing process. In this study, we have analyzed the relative binding affinities of two different gRNA/mRNA pairs. The affinity of gA6-14 for its message (ATPase 6) is high, with an apparent K(D) in the 5-10 nM range. In contrast, gCYb-558 has a low affinity for its cognate mRNA. Deletion of the gRNA U-tail caused a significant reduction in the binding affinity for only the gCYb-558 pair, and was observed only under physiological magnesium conditions. These results indicate that the U-tail contribution can differ substantially between the different gRNA/mRNA pairs. In addition, our results suggest that the efficiency of gRNA/mRNA interaction is highly dependent on thermodynamic parameters determined by the local sequences and their adopted structures surrounding the anchor-binding site.
Figures


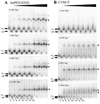
 indicate the positions of the major gRNA/mRNA complexes. For the gA6-14/3’A6PES1 pair a second complex (*) of slower mobility was observed at high mRNA concentrations. For the gA6-14/3’A6PES1 interaction, the following concentrations of pre-edited mRNAs were employed: 0 mRNA (lanes 1, 2), 1.25nM (lanes 3, 4), 2.5nM (lanes 5, 6), 5.0nM (lanes 7, 8), 12.5nM (lanes 9, 10), 25nM (lanes 11, 12), 50nM (lanes 13, 14), 100 nM (lanes 15,16). For the gCYb-558/CYbUT interaction, the following concentrations were used: 0 mRNA (lanes 1, 2), 0.5µM (lanes 3, 4), 0.75µM (lanes 5, 6), 1.0µM (lanes 7, 8), 1.5µM (lanes 9, 10), 2.0µM (lanes 11, 12), 2.25µM (lanes 13, 14), 2.5µM (lanes 15, 16).
indicate the positions of the major gRNA/mRNA complexes. For the gA6-14/3’A6PES1 pair a second complex (*) of slower mobility was observed at high mRNA concentrations. For the gA6-14/3’A6PES1 interaction, the following concentrations of pre-edited mRNAs were employed: 0 mRNA (lanes 1, 2), 1.25nM (lanes 3, 4), 2.5nM (lanes 5, 6), 5.0nM (lanes 7, 8), 12.5nM (lanes 9, 10), 25nM (lanes 11, 12), 50nM (lanes 13, 14), 100 nM (lanes 15,16). For the gCYb-558/CYbUT interaction, the following concentrations were used: 0 mRNA (lanes 1, 2), 0.5µM (lanes 3, 4), 0.75µM (lanes 5, 6), 1.0µM (lanes 7, 8), 1.5µM (lanes 9, 10), 2.0µM (lanes 11, 12), 2.25µM (lanes 13, 14), 2.5µM (lanes 15, 16).

Similar articles
-
Interactions of mRNAs and gRNAs involved in trypanosome mitochondrial RNA editing: structure probing of a gRNA bound to its cognate mRNA.RNA. 2006 Jun;12(6):1050-60. doi: 10.1261/rna.3406. Epub 2006 Apr 17. RNA. 2006. PMID: 16618968 Free PMC article.
-
RNA editing in Trypanosoma brucei: characterization of gRNA U-tail interactions with partially edited mRNA substrates.Nucleic Acids Res. 2001 Feb 1;29(3):703-9. doi: 10.1093/nar/29.3.703. Nucleic Acids Res. 2001. PMID: 11160892 Free PMC article.
-
The impact of mRNA structure on guide RNA targeting in kinetoplastid RNA editing.PLoS One. 2010 Aug 17;5(8):e12235. doi: 10.1371/journal.pone.0012235. PLoS One. 2010. PMID: 20808932 Free PMC article.
-
Mitochondrial RNA editing in trypanosomes: small RNAs in control.Biochimie. 2014 May;100:125-31. doi: 10.1016/j.biochi.2014.01.003. Epub 2014 Jan 17. Biochimie. 2014. PMID: 24440637 Free PMC article. Review.
-
The mechanism of U insertion/deletion RNA editing in kinetoplastid mitochondria.Nucleic Acids Res. 1997 Oct 1;25(19):3751-9. doi: 10.1093/nar/25.19.3751. Nucleic Acids Res. 1997. PMID: 9380494 Free PMC article. Review.
Cited by
-
Trypanosoma brucei ATPase subunit 6 mRNA bound to gA6-14 forms a conserved three-helical structure.RNA. 2008 Oct;14(10):2195-211. doi: 10.1261/rna.1144508. Epub 2008 Sep 4. RNA. 2008. PMID: 18772247 Free PMC article.
-
Interactions of mRNAs and gRNAs involved in trypanosome mitochondrial RNA editing: structure probing of a gRNA bound to its cognate mRNA.RNA. 2006 Jun;12(6):1050-60. doi: 10.1261/rna.3406. Epub 2006 Apr 17. RNA. 2006. PMID: 16618968 Free PMC article.
-
Trypanosome RNA Editing Mediator Complex proteins have distinct functions in gRNA utilization.Nucleic Acids Res. 2017 Jul 27;45(13):7965-7983. doi: 10.1093/nar/gkx458. Nucleic Acids Res. 2017. PMID: 28535252 Free PMC article.
-
The crystal structure of an oligo(U):pre-mRNA duplex from a trypanosome RNA editing substrate.RNA. 2011 Oct;17(10):1870-83. doi: 10.1261/rna.2880311. Epub 2011 Aug 30. RNA. 2011. PMID: 21878548 Free PMC article.
-
REH2 RNA helicase in kinetoplastid mitochondria: ribonucleoprotein complexes and essential motifs for unwinding and guide RNA (gRNA) binding.J Biol Chem. 2010 Jan 8;285(2):1220-8. doi: 10.1074/jbc.M109.051862. Epub 2009 Oct 22. J Biol Chem. 2010. PMID: 19850921 Free PMC article.
References
-
- Madison-Antenucci S, Grams J, Hajduk SL. Editing Machines: The complexities of Trypanosome RNA Editing. Cell. 2002;108:435–438. - PubMed
-
- Stuart K, Panigrahi AK. RNA editing: complexity and complications. Molecular Microbiology. 2002;45:591–596. - PubMed
-
- Blum B, Bakalara N, Simpson L. A model for RNA editing in kinetoplastid mitochondria: "Guide" RNA molecules transcribed from maxicircle DNA provide the edited information. Cell. 1990;60:189–198. - PubMed
-
- Blum B, Simpson L. Guide RNAs in kinetoplastid mitochondria have a nonencoded 3' oligo-(U) tail involved in recognition of the pre-edited region. Cell. 1990;62:391–397. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
