Modification and optimization of the united-residue (UNRES) potential energy function for canonical simulations. I. Temperature dependence of the effective energy function and tests of the optimization method with single training proteins
- PMID: 17201450
- PMCID: PMC3236617
- DOI: 10.1021/jp065380a
Modification and optimization of the united-residue (UNRES) potential energy function for canonical simulations. I. Temperature dependence of the effective energy function and tests of the optimization method with single training proteins
Abstract
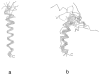
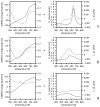
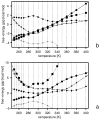
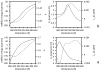
We report the modification and parametrization of the united-residue (UNRES) force field for energy-based protein structure prediction and protein folding simulations. We tested the approach on three training proteins separately: 1E0L (beta), 1GAB (alpha), and 1E0G (alpha + beta). Heretofore, the UNRES force field had been designed and parametrized to locate native-like structures of proteins as global minima of their effective potential energy surfaces, which largely neglected the conformational entropy because decoys composed of only lowest-energy conformations were used to optimize the force field. Recently, we developed a mesoscopic dynamics procedure for UNRES and applied it with success to simulate protein folding pathways. However, the force field turned out to be largely biased toward -helical structures in canonical simulations because the conformational entropy had been neglected in the parametrization. We applied the hierarchical optimization method, developed in our earlier work, to optimize the force field; in this method, the conformational space of a training protein is divided into levels, each corresponding to a certain degree of native-likeness. The levels are ordered according to increasing native-likeness; level 0 corresponds to structures with no native-like elements, and the highest level corresponds to the fully native-like structures. The aim of optimization is to achieve the order of the free energies of levels, decreasing as their native-likeness increases. The procedure is iterative, and decoys of the training protein(s) generated with the energy function parameters of the preceding iteration are used to optimize the force field in a current iteration. We applied the multiplexing replica-exchange molecular dynamics (MREMD) method, recently implemented in UNRES, to generate decoys; with this modification, conformational entropy is taken into account. Moreover, we optimized the free-energy gaps between levels at temperatures corresponding to a predominance of folded or unfolded structures, as well as to structures at the putative folding-transition temperature, changing the sign of the gaps at the transition temperature. This enabled us to obtain force fields characterized by a single peak in the heat capacity at the transition temperature. Furthermore, we introduced temperature dependence to the UNRES force field; this is consistent with the fact that it is a free-energy and not a potential energy function. beta
Figures


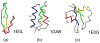











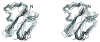





References
-
- Skolnick J, Zhang Y, Arakaki AK, Kolińsi A, Boniecki M, Szilagyi A, Kihara D. Proteins: Struct Func Genet. 2003;53:469. - PubMed
-
- Eskow E, Bader D, Byrd R, Crivelli S, Head-Gordon T, Lamberti V, Schnabel R. Math Program. 2004;101:497.
-
- Fujitsuka Y, Takada S, Luthey-Schulten ZA, Wolynes PG. Proteins: Struct Func Genet. 2004;54:88. - PubMed
-
- Scheraga HA, Liwo A, Ołdziej S, Czaplewski C, Pillardy J, Ripoll DR, Vila JA, KaŸmierkiewicz R, Saunders JA, Arnautova YA, Jagielska A, Chinchio M, Nanias M. Frontiers in Bioscience. 2004;9:3296. - PubMed
-
- Petrey D, Honig B. Mol Cell. 2005;20:811. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

