Radical sites in Mycobacterium tuberculosis KatG identified using electron paramagnetic resonance spectroscopy, the three-dimensional crystal structure, and electron transfer couplings
- PMID: 17204474
- PMCID: PMC1885898
- DOI: 10.1074/jbc.M607309200
Radical sites in Mycobacterium tuberculosis KatG identified using electron paramagnetic resonance spectroscopy, the three-dimensional crystal structure, and electron transfer couplings
Abstract
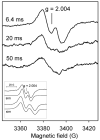
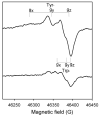
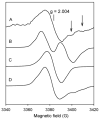
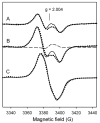
Catalase-peroxidase (KatG) from Mycobacterium tuberculosis, a Class I peroxidase, exhibits high catalase activity and peroxidase activity with various substrates and is responsible for activation of the commonly used antitubercular drug, isoniazid (INH). KatG readily forms amino acid-based radicals during turnover with alkyl peroxides, and this work focuses on extending the identification and characterization of radicals forming on the millisecond to second time scale. Rapid freeze-quench electron paramagnetic resonance spectroscopy (RFQ-EPR) reveals a change in the structure of the initially formed radical in the presence of INH. Heme pocket binding of the drug and knowledge that KatG[Y229F] lacks this signal provides evidence for radical formation on residue Tyr(229). High field RFQ-EPR spectroscopy confirmed a tryptophanyl radical signal, and new analyses of X-band RFQ-EPR spectra also established its presence. High field EPR spectroscopy also confirmed that the majority radical species is a tyrosyl radical. Site-directed mutagenesis, along with simulations of EPR spectra based on x-ray structural data for particular tyrosine and tryptophan residues, enabled assignments based on predicted hyperfine coupling parameters. KatG mutants W107F, Y229F, and the double mutant W107F/Y229F showed alteration in type and yield of radical species. Results are consistent with formation of a tyrosyl radical reasonably assigned to residue Tyr(229) within the first few milliseconds of turnover. This is followed by a mixture of tyrosyl and tryptophanyl radical species and finally to only a tyrosyl radical on residue Tyr(353), which lies more distant from the heme. The radical processing of enzyme lacking the Trp(107)-Tyr(229)-Met(255) adduct (found as a unique structural feature of catalase-peroxidases) is suggested to be a reasonable assignment of the phenomena.
Figures


Similar articles
-
Rapid formation of compound II and a tyrosyl radical in the Y229F mutant of Mycobacterium tuberculosis catalase-peroxidase disrupts catalase but not peroxidase function.J Biol Chem. 2003 Nov 7;278(45):44121-7. doi: 10.1074/jbc.M304757200. Epub 2003 Aug 27. J Biol Chem. 2003. PMID: 12944408
-
Identification and characterization of tyrosyl radical formation in Mycobacterium tuberculosis catalase-peroxidase (KatG).J Biol Chem. 2002 Nov 8;277(45):42633-8. doi: 10.1074/jbc.M207916200. Epub 2002 Aug 29. J Biol Chem. 2002. PMID: 12205099
-
Evidence for radical formation at Tyr-353 in Mycobacterium tuberculosis catalase-peroxidase (KatG).J Biol Chem. 2004 Feb 27;279(9):7606-12. doi: 10.1074/jbc.M311884200. Epub 2003 Dec 9. J Biol Chem. 2004. PMID: 14665627
-
Catalase in peroxidase clothing: Interdependent cooperation of two cofactors in the catalytic versatility of KatG.Arch Biochem Biophys. 2014 Feb 15;544:27-39. doi: 10.1016/j.abb.2013.11.007. Epub 2013 Nov 23. Arch Biochem Biophys. 2014. PMID: 24280274 Review.
-
Structure and interactions of amino acid radicals in class I ribonucleotide reductase studied by ENDOR and high-field EPR spectroscopy.Biochim Biophys Acta. 2005 Feb 25;1707(1):67-90. doi: 10.1016/j.bbabio.2004.02.011. Biochim Biophys Acta. 2005. PMID: 15721607 Review.
Cited by
-
Observation of organometallic and radical intermediates formed during the reaction of methyl-coenzyme M reductase with bromoethanesulfonate.Biochemistry. 2010 Aug 17;49(32):6866-76. doi: 10.1021/bi100650m. Biochemistry. 2010. PMID: 20597483 Free PMC article.
-
Antibiotic resistance in Mycobacterium tuberculosis: peroxidase intermediate bypass causes poor isoniazid activation by the S315G mutant of M. tuberculosis catalase-peroxidase (KatG).J Biol Chem. 2009 Jun 12;284(24):16146-16155. doi: 10.1074/jbc.M109.005546. Epub 2009 Apr 9. J Biol Chem. 2009. PMID: 19363028 Free PMC article.
-
Catalase-peroxidase (KatG): a potential frontier in tuberculosis drug development.Crit Rev Biochem Mol Biol. 2024 Dec;59(6):434-446. doi: 10.1080/10409238.2025.2470630. Epub 2025 Feb 27. Crit Rev Biochem Mol Biol. 2024. PMID: 40013498 Review.
-
Spin trapping investigation of peroxide- and isoniazid-induced radicals in Mycobacterium tuberculosis catalase-peroxidase.Biochemistry. 2008 Oct 28;47(43):11377-85. doi: 10.1021/bi800952b. Epub 2008 Oct 2. Biochemistry. 2008. PMID: 18831539 Free PMC article.
-
Two tyrosyl radicals stabilize high oxidation states in cytochrome C oxidase for efficient energy conservation and proton translocation.J Am Chem Soc. 2012 Mar 14;134(10):4753-61. doi: 10.1021/ja210535w. Epub 2012 Mar 6. J Am Chem Soc. 2012. PMID: 22296274 Free PMC article.
References
-
- Rouse DA, DeVito JA, Li Z, Byer H, Morris SL. Mol Microbiol. 1996;22 (3):583–592. - PubMed
-
- Jakopitsch C, Droghetti E, Schmuckenschlager F, Furtmuller PG, Smulevich G, Obinger C. J Biol Chem. 2005;280(51):42411–42422. - PubMed
-
- Smulevich G, Jakopitsch C, Droghetti E, Obinger C. J Inorg Biochem. 2006;100(4):568–585. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

