Particle simulation approach for subcellular dynamics and interactions of biological molecules
- PMID: 17217513
- PMCID: PMC1780110
- DOI: 10.1186/1471-2105-7-S4-S20
Particle simulation approach for subcellular dynamics and interactions of biological molecules
Abstract
Background: Spatio-temporal dynamics within cells can now be visualized at appropriate resolution, due to the advances in molecular imaging technologies. Even single-particle tracking (SPT) and single fluorophore video imaging (SFVI) are now being applied to observation of molecular-level dynamics. However, little is known concerning how molecular-level dynamics affect properties at the cellular level.
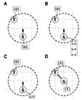
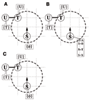
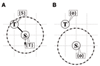
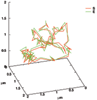
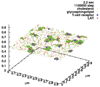
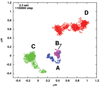
Results: We propose an algorithm designed for three-dimensional simulation of the reaction-diffusion dynamics of molecules, based on a particle model. Chemical reactions proceed through the interactions of particles in space, with activation energies determining the rates of these chemical reactions at each interaction. This energy-based model can include the cellular membrane, membranes of other organelles, and cytoskeleton. The simulation algorithm was tested for a reversible enzyme reaction model and its validity was confirmed. Snapshot images taken from simulated molecular interactions on the cell-surface revealed clustering domains (size approximately 0.2 microm) associated with rafts. Sample trajectories of raft constructs exhibited "hop diffusion". These domains corralled the diffusive motion of membrane proteins.
Conclusion: These findings demonstrate that our approach is promising for modelling the localization properties of biological phenomena.
Figures


Similar articles
-
Single particle tracking of complex diffusion in membranes: simulation and detection of barrier, raft, and interaction phenomena.J Phys Chem B. 2007 Apr 12;111(14):3625-32. doi: 10.1021/jp067187m. Epub 2007 Mar 17. J Phys Chem B. 2007. PMID: 17388520
-
Brownian dynamics simulations with hard-body interactions: spherical particles.J Chem Phys. 2012 Oct 28;137(16):164108. doi: 10.1063/1.4761827. J Chem Phys. 2012. PMID: 23126696
-
ProtNet: a tool for stochastic simulations of protein interaction networks dynamics.BMC Bioinformatics. 2007 Mar 8;8 Suppl 1(Suppl 1):S4. doi: 10.1186/1471-2105-8-S1-S4. BMC Bioinformatics. 2007. PMID: 17430571 Free PMC article.
-
Fluorescence correlation spectroscopy: linking molecular dynamics to biological function in vitro and in situ.Curr Opin Struct Biol. 2011 Oct;21(5):650-60. doi: 10.1016/j.sbi.2011.06.006. Epub 2011 Jul 21. Curr Opin Struct Biol. 2011. PMID: 21767945 Review.
-
Quantitative microscopy: protein dynamics and membrane organisation.Traffic. 2009 Aug;10(8):962-71. doi: 10.1111/j.1600-0854.2009.00908.x. Epub 2009 Apr 4. Traffic. 2009. PMID: 19416480 Review.
Cited by
-
Development of computations in bioscience and bioinformatics and its application: review of the Symposium of Computations in Bioinformatics and Bioscience (SCBB06).BMC Bioinformatics. 2006 Dec 12;7 Suppl 4(Suppl 4):S1. doi: 10.1186/1471-2105-7-S4-S1. BMC Bioinformatics. 2006. PMID: 17217501 Free PMC article.
-
Facing glycosphingolipid-Shiga toxin interaction: dire straits for endothelial cells of the human vasculature.Cell Mol Life Sci. 2013 Feb;70(3):425-57. doi: 10.1007/s00018-012-1060-z. Epub 2012 Jul 6. Cell Mol Life Sci. 2013. PMID: 22766973 Free PMC article. Review.
-
Expression of excess receptors and negative feedback control of signal pathways are required for rapid activation and prompt cessation of signal transduction.Cell Commun Signal. 2009 Mar 3;7:3. doi: 10.1186/1478-811X-7-3. Cell Commun Signal. 2009. PMID: 19254388 Free PMC article.
-
Protein charge and mass contribute to the spatio-temporal dynamics of protein-protein interactions in a minimal proteome.Proteomics. 2013 Apr;13(8):1339-51. doi: 10.1002/pmic.201100540. Epub 2013 Mar 18. Proteomics. 2013. PMID: 23420643 Free PMC article.
-
STEPS: efficient simulation of stochastic reaction-diffusion models in realistic morphologies.BMC Syst Biol. 2012 May 10;6:36. doi: 10.1186/1752-0509-6-36. BMC Syst Biol. 2012. PMID: 22574658 Free PMC article.
References
-
- Ritchie K, Kusumi A. Single-particle tracking image microscopy. Methods Enzymol. 2003;360:618–34. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

