Modes and modulations of antibiotic resistance gene expression
- PMID: 17223624
- PMCID: PMC1797629
- DOI: 10.1128/CMR.00015-06
Modes and modulations of antibiotic resistance gene expression
Abstract
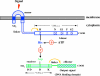
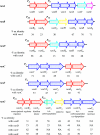
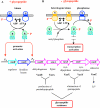
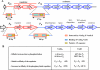
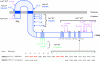
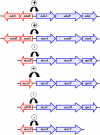
Since antibiotic resistance usually affords a gain of function, there is an associated biological cost resulting in a loss of fitness of the bacterial host. Considering that antibiotic resistance is most often only transiently advantageous to bacteria, an efficient and elegant way for them to escape the lethal action of drugs is the alteration of resistance gene expression. It appears that expression of bacterial resistance to antibiotics is frequently regulated, which indicates that modulation of gene expression probably reflects a good compromise between energy saving and adjustment to a rapidly evolving environment. Modulation of gene expression can occur at the transcriptional or translational level following mutations or the movement of mobile genetic elements and may involve induction by the antibiotic. In the latter case, the antibiotic can have a triple activity: as an antibacterial agent, as an inducer of resistance to itself, and as an inducer of the dissemination of resistance determinants. We will review certain mechanisms, all reversible, that bacteria have elaborated to achieve antibiotic resistance by the fine-tuning of the expression of genetic information.
Figures
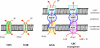
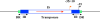



References
-
- Aendekerk, S., B. Ghysels, P. Cornelis, and C. Baysse. 2002. Characterization of a new efflux pump, MexGHI-OpmD, from Pseudomonas aeruginosa that confers resistance to vanadium. Microbiology 148:2371-2381. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

