Interaction of yeast RNA-binding proteins Nrd1 and Nab3 with RNA polymerase II terminator elements
- PMID: 17237360
- PMCID: PMC1800511
- DOI: 10.1261/rna.338407
Interaction of yeast RNA-binding proteins Nrd1 and Nab3 with RNA polymerase II terminator elements
Abstract
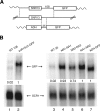
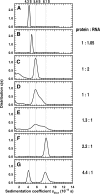
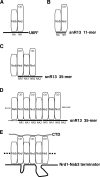
Yeast RNA-binding proteins Nrd1 and Nab3 direct transcription termination of sn/snoRNA transcripts, some mRNA transcripts, and a class of intergenic and anti-sense transcripts. Recognition of Nrd1- and Nab3-binding sites is a critical first step in the termination and subsequent processing or degradation of these transcripts. In this article, we describe the purification and characterization of an Nrd1-Nab3 heterodimer. This Nrd1-Nab3 complex binds specifically to RNA sequences derived from a snoRNA terminator. The relative binding to mutant terminators correlates with the in vivo termination efficiency of these mutations, indicating that the primary specificity determinant in nonpoly(A) termination is Nrd1-Nab3 binding. In addition, several snoRNA terminators contain multiple Nrd1- and Nab3-binding sites and we show that multiple heterodimers bind cooperatively to one of these terminators in vitro.
Figures




Similar articles
-
Identification of cis elements directing termination of yeast nonpolyadenylated snoRNA transcripts.Mol Cell Biol. 2004 Jul;24(14):6241-52. doi: 10.1128/MCB.24.14.6241-6252.2004. Mol Cell Biol. 2004. PMID: 15226427 Free PMC article.
-
Regulation of yeast NRD1 expression by premature transcription termination.Mol Cell. 2006 Mar 3;21(5):641-51. doi: 10.1016/j.molcel.2006.02.005. Mol Cell. 2006. PMID: 16507362
-
Termination of cryptic unstable transcripts is directed by yeast RNA-binding proteins Nrd1 and Nab3.Mol Cell. 2006 Sep 15;23(6):841-51. doi: 10.1016/j.molcel.2006.07.024. Mol Cell. 2006. PMID: 16973436
-
The Nrd1-Nab3-Sen1 transcription termination complex from a structural perspective.Biochem Soc Trans. 2023 Jun 28;51(3):1257-1269. doi: 10.1042/BST20221418. Biochem Soc Trans. 2023. PMID: 37222282 Free PMC article. Review.
-
Termination of Transcription of Short Noncoding RNAs by RNA Polymerase II.Annu Rev Biochem. 2015;84:381-404. doi: 10.1146/annurev-biochem-060614-034457. Epub 2015 Mar 26. Annu Rev Biochem. 2015. PMID: 25747400 Free PMC article. Review.
Cited by
-
The Saccharomyces cerevisiae Nrd1-Nab3 transcription termination pathway acts in opposition to Ras signaling and mediates response to nutrient depletion.Mol Cell Biol. 2012 May;32(10):1762-75. doi: 10.1128/MCB.00050-12. Epub 2012 Mar 19. Mol Cell Biol. 2012. PMID: 22431520 Free PMC article.
-
Purification and In Vitro Analysis of the Exosome Cofactors Nrd1-Nab3 and Trf4-Air2.Methods Mol Biol. 2020;2062:277-289. doi: 10.1007/978-1-4939-9822-7_14. Methods Mol Biol. 2020. PMID: 31768982
-
Writing a wrong: Coupled RNA polymerase II transcription and RNA quality control.Wiley Interdiscip Rev RNA. 2019 Jul;10(4):e1529. doi: 10.1002/wrna.1529. Epub 2019 Mar 7. Wiley Interdiscip Rev RNA. 2019. PMID: 30848101 Free PMC article. Review.
-
A network of interdependent molecular interactions describes a higher order Nrd1-Nab3 complex involved in yeast transcription termination.J Biol Chem. 2013 Nov 22;288(47):34158-34167. doi: 10.1074/jbc.M113.516765. Epub 2013 Oct 7. J Biol Chem. 2013. PMID: 24100036 Free PMC article.
-
Recombinant protein complex expression in E. coli.Curr Protoc Protein Sci. 2008 May;Chapter 5:Unit 5.21. doi: 10.1002/0471140864.ps0521s52. Curr Protoc Protein Sci. 2008. PMID: 18491300 Free PMC article.
References
-
- Arigo, J.T., Carroll, K.L., Ames, J.M., Corden, J.L. Regulation of yeast NRD1 expression by premature transcription termination. Mol. Cell. 2006a;21:641–651. - PubMed
-
- Arigo, J.T., Eyler, D.E., Carroll, K.L., Corden, J.L. Termination of cryptic unstable transcripts is directed by yeast RNA-binding proteins Nrd1 and Nab3. Mol. Cell. 2006b;23:841–851. - PubMed
-
- Bentley, D.L. Rules of engagement: Co-transcriptional recruitment of pre-mRNA processing factors. Curr. Opin. Cell Biol. 2005;17:251–256. - PubMed
-
- Brachmann, C.B., Davies, A., Cost, G.J., Caputo, E., Li, J., Hieter, P., Boeke, J.D. Designer deletion strains derived from Saccharomyces cerevisiae S288C: A useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast. 1998;14:115–132. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
