Cytosolic and ER J-domains of mammalian and parasitic origin can functionally interact with DnaK
- PMID: 17239655
- PMCID: PMC1906734
- DOI: 10.1016/j.biocel.2006.11.006
Cytosolic and ER J-domains of mammalian and parasitic origin can functionally interact with DnaK
Abstract
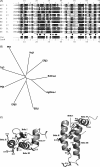
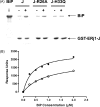
Both prokaryotic and eukaryotic cells contain multiple heat shock protein 40 (Hsp40) and heat shock protein 70 (Hsp70) proteins, which cooperate as molecular chaperones to ensure fidelity at all stages of protein biogenesis. The Hsp40 signature domain, the J-domain, is required for binding of an Hsp40 to a partner Hsp70, and may also play a role in the specificity of the association. Through the creation of chimeric Hsp40 proteins by the replacement of the J-domain of a prokaryotic Hsp40 (DnaJ), we have tested the functional equivalence of J-domains from a number of divergent Hsp40s of mammalian and parasitic origin (malarial Pfj1 and Pfj4, trypanosomal Tcj3, human ERj3, ERj5, and Hsj1, and murine ERj1). An in vivo functional assay was used to test the functionality of the chimeric proteins on the basis of their ability to reverse the thermosensitivity of a dnaJ cbpA mutant Escherichia coli strain (OD259). The Hsp40 chimeras containing J-domains originating from soluble (cytosolic or endoplasmic reticulum (ER)-lumenal) Hsp40s were able to reverse the thermosensitivity of E. coli OD259. In all cases, modified derivatives of these chimeric proteins containing an His to Gln substitution in the HPD motif of the J-domain were unable to reverse the thermosensitivity of E. coli OD259. This suggested that these J-domains exerted their in vivo functionality through a specific interaction with E. coli Hsp70, DnaK. Interestingly, a Hsp40 chimera containing the J-domain of ERj1, an integral membrane-bound ER Hsp40, was unable to reverse the thermosensitivity of E. coli OD259, suggesting that this J-domain was unable to functionally interact with DnaK. Substitutions of conserved amino acid residues and motifs were made in all four helices (I-IV) and the loop regions of the J-domains, and the modified chimeric Hsp40s were tested for functionality using the in vivo assay. Substitution of a highly conserved basic residue in helix II of the J-domain was found to disrupt in vivo functionality for all the J-domains tested. We propose that helix II and the HPD motif of the J-domain represent the fundamental elements of a binding surface required for the interaction of Hsp40s with Hsp70s, and that this surface has been conserved in mammalian, parasitic and bacterial systems.
Figures


Similar articles
-
Complementation Assays for Co-chaperone Function.Methods Mol Biol. 2023;2693:105-111. doi: 10.1007/978-1-0716-3342-7_9. Methods Mol Biol. 2023. PMID: 37540430
-
Rational mutagenesis of a 40 kDa heat shock protein from Agrobacterium tumefaciens identifies amino acid residues critical to its in vivo function.Int J Biochem Cell Biol. 2005 Jan;37(1):177-91. doi: 10.1016/j.biocel.2004.06.009. Int J Biochem Cell Biol. 2005. PMID: 15381160
-
Structure and energetics of an allele-specific genetic interaction between dnaJ and dnaK: correlation of nuclear magnetic resonance chemical shift perturbations in the J-domain of Hsp40/DnaJ with binding affinity for the ATPase domain of Hsp70/DnaK.Biochemistry. 2003 May 6;42(17):4926-36. doi: 10.1021/bi027070y. Biochemistry. 2003. PMID: 12718534
-
The diversity of the DnaJ/Hsp40 family, the crucial partners for Hsp70 chaperones.Cell Mol Life Sci. 2006 Nov;63(22):2560-70. doi: 10.1007/s00018-006-6192-6. Cell Mol Life Sci. 2006. PMID: 16952052 Free PMC article. Review.
-
Not all J domains are created equal: implications for the specificity of Hsp40-Hsp70 interactions.Protein Sci. 2005 Jul;14(7):1697-709. doi: 10.1110/ps.051406805. Protein Sci. 2005. PMID: 15987899 Free PMC article. Review.
Cited by
-
Plasmodium falciparum Molecular Chaperones: Guardians of the Malaria Parasite Proteome and Renovators of the Host Proteome.Front Cell Dev Biol. 2022 May 16;10:921739. doi: 10.3389/fcell.2022.921739. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35652103 Free PMC article. Review.
-
Functional relevance of J-protein family of rice (Oryza sativa).Cell Stress Chaperones. 2013 May;18(3):321-31. doi: 10.1007/s12192-012-0384-9. Epub 2012 Nov 16. Cell Stress Chaperones. 2013. PMID: 23160806 Free PMC article.
-
The structural and functional diversity of Hsp70 proteins from Plasmodium falciparum.Protein Sci. 2007 Sep;16(9):1803-18. doi: 10.1110/ps.072918107. Protein Sci. 2007. PMID: 17766381 Free PMC article. Review.
-
Plasmodium falciparum encodes a single cytosolic type I Hsp40 that functionally interacts with Hsp70 and is upregulated by heat shock.Cell Stress Chaperones. 2011 Jul;16(4):389-401. doi: 10.1007/s12192-010-0250-6. Epub 2010 Dec 30. Cell Stress Chaperones. 2011. PMID: 21191678 Free PMC article.
-
In silico identification of modulators of J domain protein-Hsp70 interactions in Plasmodium falciparum: a drug repurposing strategy against malaria.Front Mol Biosci. 2023 Aug 9;10:1158912. doi: 10.3389/fmolb.2023.1158912. eCollection 2023. Front Mol Biosci. 2023. PMID: 37621993 Free PMC article.
References
-
- Berjanskii M.V., Riley M.I., Xie A., Semenchenko C., Folk W.R., Van Doren S.R. NMR structure of the N-terminal J domain of murine polyomavirus T antigens. Implications for DnaJ-like domains and for mutations of T antigens. The Journal of Biological Chemistry. 2000;275:36094–36103. - PubMed
-
- Bies C., Guth S., Janoschek K., Nastainczyk W., Volkmer J., Zimmermann R. A Scj1p homologue and folding catalyst present in Dog pancreas microsomes. Biological Chemistry. 1999;380:1175–1182. - PubMed
-
- Bies C., Blum R., Dudek J., Nastainczyk W., Oberhauser S., Jung M. Characterization of pancreatic ERj3p, a homolog of yeast DnaJ-like protein Scj1p. Biological Chemistry. 2004;385:389–395. - PubMed
-
- Brightman S.J., Blatch G.L., Zetter B.R. Isolation of a mouse cDNA encoding MTJ1, a new murine member of the DnaJ family of proteins. Gene. 1995;153:249–254. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

