Inverted DNA repeats channel repair of distant double-strand breaks into chromatid fusions and chromosomal rearrangements
- PMID: 17242181
- PMCID: PMC1899885
- DOI: 10.1128/MCB.01740-06
Inverted DNA repeats channel repair of distant double-strand breaks into chromatid fusions and chromosomal rearrangements
Abstract
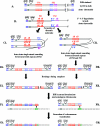
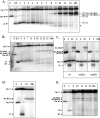
Inverted DNA repeats are known to cause genomic instabilities. Here we demonstrate that double-strand DNA breaks (DSBs) introduced a large distance from inverted repeats in the yeast (Saccharomyces cerevisiae) chromosome lead to a burst of genomic instability. Inverted repeats located as far as 21 kb from each other caused chromosome rearrangements in response to a single DSB. We demonstrate that the DSB initiates a pairing interaction between inverted repeats, resulting in the formation of large dicentric inverted dimers. Furthermore, we observed that propagation of cells containing inverted dimers led to gross chromosomal rearrangements, including translocations, truncations, and amplifications. Finally, our data suggest that break-induced replication is responsible for the formation of translocations resulting from anaphase breakage of inverted dimers. We propose a model explaining the formation of inverted dicentric dimers by intermolecular single-strand annealing (SSA) between inverted DNA repeats. According to this model, anaphase breakage of inverted dicentric dimers leads to gross chromosomal rearrangements (GCR). This "SSA-GCR" pathway is likely to be important in the repair of isochromatid breaks resulting from collapsed replication forks, certain types of radiation, or telomere aberrations that mimic isochromatid breaks.
Figures

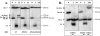



Similar articles
-
Large inverted repeats in the vicinity of a single double-strand break strongly affect repair in yeast diploids lacking Rad51.Mutat Res. 2008 Oct 14;645(1-2):9-18. doi: 10.1016/j.mrfmmm.2008.07.013. Epub 2008 Aug 5. Mutat Res. 2008. PMID: 18755201 Free PMC article.
-
Fusion of nearby inverted repeats by a replication-based mechanism leads to formation of dicentric and acentric chromosomes that cause genome instability in budding yeast.Genes Dev. 2009 Dec 15;23(24):2861-75. doi: 10.1101/gad.1862709. Genes Dev. 2009. PMID: 20008936 Free PMC article.
-
Single-strand annealing between inverted DNA repeats: Pathway choice, participating proteins, and genome destabilizing consequences.PLoS Genet. 2018 Aug 9;14(8):e1007543. doi: 10.1371/journal.pgen.1007543. eCollection 2018 Aug. PLoS Genet. 2018. PMID: 30091972 Free PMC article.
-
Repair and chromosomal damage.Radiother Oncol. 2004 Sep;72(3):251-6. doi: 10.1016/j.radonc.2004.07.005. Radiother Oncol. 2004. PMID: 15450722 Review.
-
Faithful after break-up: suppression of chromosomal translocations.Cell Mol Life Sci. 2009 Oct;66(19):3149-60. doi: 10.1007/s00018-009-0068-5. Epub 2009 Jun 23. Cell Mol Life Sci. 2009. PMID: 19547915 Free PMC article. Review.
Cited by
-
Break-induced replication: functions and molecular mechanism.Curr Opin Genet Dev. 2013 Jun;23(3):271-9. doi: 10.1016/j.gde.2013.05.007. Epub 2013 Jun 18. Curr Opin Genet Dev. 2013. PMID: 23790415 Free PMC article. Review.
-
Defective break-induced replication leads to half-crossovers in Saccharomyces cerevisiae.Genetics. 2008 Aug;179(4):1845-60. doi: 10.1534/genetics.108.087940. Epub 2008 Aug 9. Genetics. 2008. PMID: 18689895 Free PMC article.
-
Chromosome position determines the success of double-strand break repair.Proc Natl Acad Sci U S A. 2016 Jan 12;113(2):E146-54. doi: 10.1073/pnas.1523660113. Epub 2015 Dec 29. Proc Natl Acad Sci U S A. 2016. PMID: 26715752 Free PMC article.
-
Extensive DNA end processing by exo1 and sgs1 inhibits break-induced replication.PLoS Genet. 2010 Jul 8;6(7):e1001007. doi: 10.1371/journal.pgen.1001007. PLoS Genet. 2010. PMID: 20628570 Free PMC article.
-
Repair of DNA Breaks by Break-Induced Replication.Annu Rev Biochem. 2021 Jun 20;90:165-191. doi: 10.1146/annurev-biochem-081420-095551. Epub 2021 Apr 1. Annu Rev Biochem. 2021. PMID: 33792375 Free PMC article. Review.
References
-
- Artandi, S. E., S. Chang, S.-L. Lee, S. Alson, and G. L. Gottlieb. 2000. Telomere dysfunction promotes non-reciprocal translocations and epithelial cancers in mice. Nature 406:641-645. - PubMed
-
- Behr, T., M. Behe, M. Stabin, E. Wehrmann, C. Apostolidis, R. Molinet, F. Strutz, A. Fayyazi, E. Wieland, S. Gratz, L. Koch, D. Goldenberg, and W. Becker. 1999. High-linear energy transfer (LET) α versus low-LET β emitters in radioimmunotherapy of solid tumors: therapeutic efficacy and dose-limiting toxicity of 213 Bi-versus 90 Y-labeled CO17-1A Fab fragments in a human colonic cancer model. Cancer Res. 59:2635-2643. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
