Archaeal MCM has separable processivity, substrate choice and helicase domains
- PMID: 17259218
- PMCID: PMC1807962
- DOI: 10.1093/nar/gkl1117
Archaeal MCM has separable processivity, substrate choice and helicase domains
Abstract
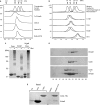
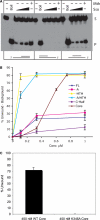
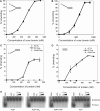
The mini-chromosome maintenance (MCM) complex is the principal candidate for the replicative helicase of archaea and eukaryotes. Here, we describe a functional dissection of the roles of the three principal structural modules of the homomultimeric MCM of the hyperthermophilic archaeon Sulfolobus solfataricus. Our results include the first analysis of the central AAA+ domain in isolation. This domain possesses ATPase and helicase activity, defining this as the minimal helicase domain. Reconstitution experiments show that the helicase activity of the AAA+ domain can be stimulated by addition of the isolated N-terminal half in trans. Addition of the N-terminus influences both the processivity of the helicase and the choice of substrate that can be melted by the ATPase domain. The degenerate helix-turn-helix domain at the C-terminus of MCM exerts a negative effect on the helicase activity of the complex. These results provide the first evidence for extensive regulatory inter-domain communication within the MCM complex.
Figures


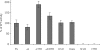

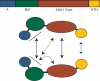
References
-
- Kelman LM, Kelman Z. Archaea: an archetype for replication initiation studies? Mol Micro. 2003;48:605–616. - PubMed
-
- Duggin IG, Bell SD. The chromosome replication machinery of the Archaeon Sulfolobus solfataricus. J. Biol. Chem. 2006;281:15029–15032. - PubMed
-
- Tye BK. MCM proteins in DNA replication. Ann. Rev. Bioch. 1999;68:649–686. - PubMed
-
- Malorano D, Cuvier O, Danis E, Mechali M. MCM8 is an MCM2-7-related protein that functions as a DNA helicase during replication elongation and not initiation. Cell. 2005;120:315–328. - PubMed

